MtoZ Biolabs provides TMT-based Quantitative Proteomics Analysis Service using isobaric tandem mass tag labeling combined with high-resolution LC-MS/MS to achieve accurate multiplexed protein quantification across multiple samples in a single run. This service supports discovery proteomics, biomarker screening, and mechanism studies across cells, tissues, and biofluids in a robust and scalable workflow.
What Is TMT-Based Quantitative Proteomics?
TMT-based quantitative proteomics is a multiplexed MS-based strategy that labels peptides from different samples with isobaric tandem mass tags, allowing them to be mixed and analyzed in a single LC-MS/MS run. At the MS1 level all labeled peptides appear with the same mass, while at the MS2 level the tags release distinct reporter ions whose intensities reflect relative peptide and protein abundance across samples.
Each TMT reagent contains three functional parts:
- Amine Reactive Group: Covalently reacts with peptide N-termini and lysine side chains, ensuring stable labeling of most peptides.
- Mass Normalization Group: Balances the total mass of different tags so that all labeled peptides are isobaric at the precursor level.
- Reporter Ion Group: Generates low-mass reporter ions with channel-specific m/z during fragmentation, providing the quantitative readout for each sample in the multiplex set.
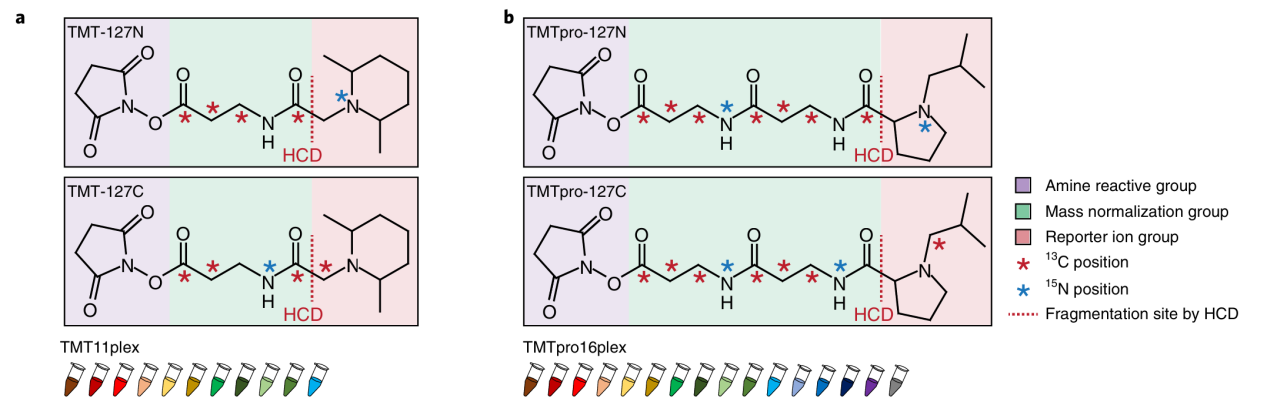
Li, J. et al. Nat Methods. 2020.
Figure 1. Chemical Structures of TMT (a) and TMTpro Reagents (b)
TMT-Based Quantitative Proteomics Analysis Service at MtoZ Biolabs
MtoZ Biolabs' TMT-based Quantitative Proteomics Analysis Service is built on Thermo Fisher Orbitrap Fusion Lumos mass spectrometers, coupled with nano-liquid chromatography and optimized TMT and TMTpro workflows. The Orbitrap Fusion Lumos platform provides ultra-high resolution and sensitivity, enabling confident identification and quantification of low-abundance peptides, while combined HCD and ETD fragmentation preserves peptide sequence information and supports accurate reporter ion–based quantitation. By labeling peptides from multiple samples with distinct TMT channels and analyzing them in a single LC-MS/MS run, we deliver high-plex comparative proteomics with deep proteome coverage, reduced technical variability, and robust detection of signaling, structural, metabolic, and regulatory proteins across complex experimental designs such as multi-group studies, time course experiments, and clinical cohorts. You only need to submit your samples, and MtoZ Biolabs manages protein extraction, digestion, TMT labeling, pooling, LC-MS/MS acquisition, data processing, and bioinformatics analysis.
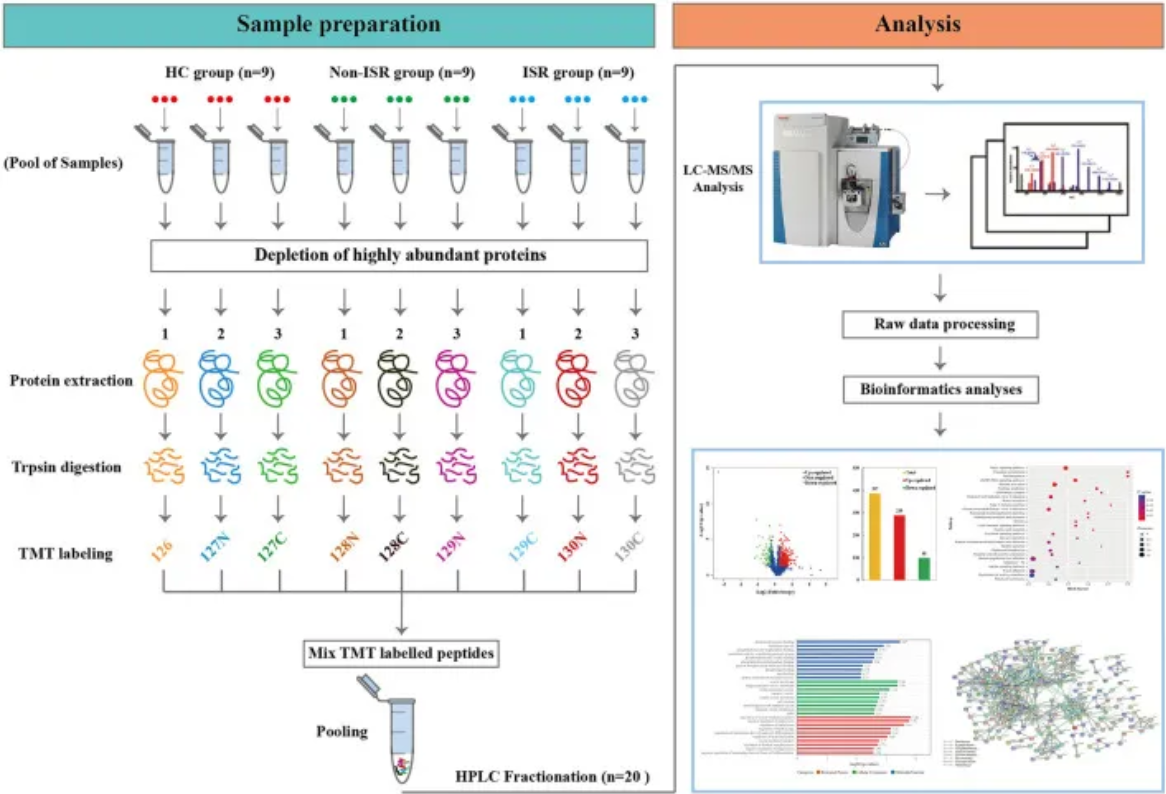
Hou, J. et al. Front Cardiovasc Med. 2022.
Figure 2. Workflow for TMT-based Quantitative Proteomics Analysis Service
Why Choose MtoZ Biolabs
1. High-Performance Platform
Built on Thermo Fisher Orbitrap Fusion Lumos with nano-LC and optimized TMT/TMTpro workflows for high resolution and sensitivity.
2. Reliable Multiplex Quantification
Accurate reporter ion–based quantification across multiple TMT channels, reducing run-to-run variation.
3. Deep Proteome Coverage
Optional peptide fractionation and refined MS methods support large numbers of quantified proteins, including low-abundance regulators.
4. Robust Data Processing
Standardized pipelines for identification, normalization, differential analysis, and functional enrichment.
5. Flexible Study Design
Supports multi-group, time course, and clinical cohort studies with clear channel assignment strategies.
6. End-to-End Support
From experimental design to data interpretation, an experienced proteomics team assists throughout your project.
Applications of TMT-Based Quantitative Proteomics Analysis Service
TMT-based quantitative proteomics at MtoZ Biolabs can be applied to many research and development scenarios.
1. Disease Versus Control Comparisons
Identify proteins that differ between healthy and diseased samples in oncology, neurology, immunology, and other fields
2. Biomarker Discovery and Prioritization
Generate candidate protein markers for diagnosis, prognosis, or treatment response
3. Drug Mechanism and Pharmacodynamics
Characterize proteomic changes in response to small molecules, biologics, or gene therapies
4. Time Course and Dose Response Studies
Track dynamic protein expression changes over time or across multiple doses in a single multiplexed design
FAQ
Q1: What types of samples are suitable?
We support a broad range of samples for TMT-based quantitative proteomics, including but not limited to:
-
Cell pellets from adherent or suspension cell lines
-
Primary cells from blood, bone marrow, or tissues
-
Fresh frozen tissues or biopsies
-
Biofluids such as serum, plasma, CSF, and urine
-
Preclinical or animal model samples for comparative studies
Q2: How should I prepare my samples?
- Provide fresh or snap-frozen samples to ensure protein integrity.
- Avoid detergents, salts, and inhibitors that interfere with TMT labeling or LC-MS/MS analysis.
- Store samples at –80 °C before shipment.
- Ship frozen samples on dry ice and clearly label sample groups for channel assignment.
Detailed requirements and recommendations can be found in the Sample Submission Guidelines for Proteomics.
Q3: What is the service general workflow?
Q4: What data formats are provided?
- Raw LC-MS/MS files in instrument specific formats.
- Peptide and protein identification and TMT reporter ion quantification tables in .xlsx or .csv.
- Statistical and functional analysis outputs (for example differential expression and pathway enrichment) in .xlsx or .csv.
- Publication ready figures such as heatmaps, volcano plots, and PCA style plots in high resolution PNG or TIFF.
- A concise PDF report summarizing experimental design, TMT setup, methods, QC metrics, and key findings.
- Additional formats can be provided on request to meet specific analysis or publication needs.
Start Your Project with MtoZ Biolabs
Our TMT-based Quantitative Proteomics Analysis Service is designed to provide more rapid, high-throughput, and cost-effective analysis, with exceptional data quality and minimal sample consumption. Free project evaluation, welcome to learn more details.
