MtoZ Biolabs employs high-resolution mass spectrometry analysis platforms to deliver tumor proteomics solutions which enable the systematic characterization of protein composition and expression profiles in tumor samples. This solution is suitable for tumor proteome profiling and comparative analysis across samples, providing reliable data support for tumor-related proteomics research.
Overview
Tumor proteomics is a research field that focuses on tumor-related samples and aims to systematically analyze protein composition, expression levels, and dynamic changes. Tumors are inherently complex diseases caused by abnormal cellular proliferation and dysregulated molecular control, and their initiation and progression are directly reflected in alterations in protein expression, post-translational modifications, and functional states. Through tumor proteomics analysis, the molecular characteristics and regulatory mechanisms of tumors can be elucidated at the protein level, with broad applications in biomarker discovery, pathway analysis, and drug development.
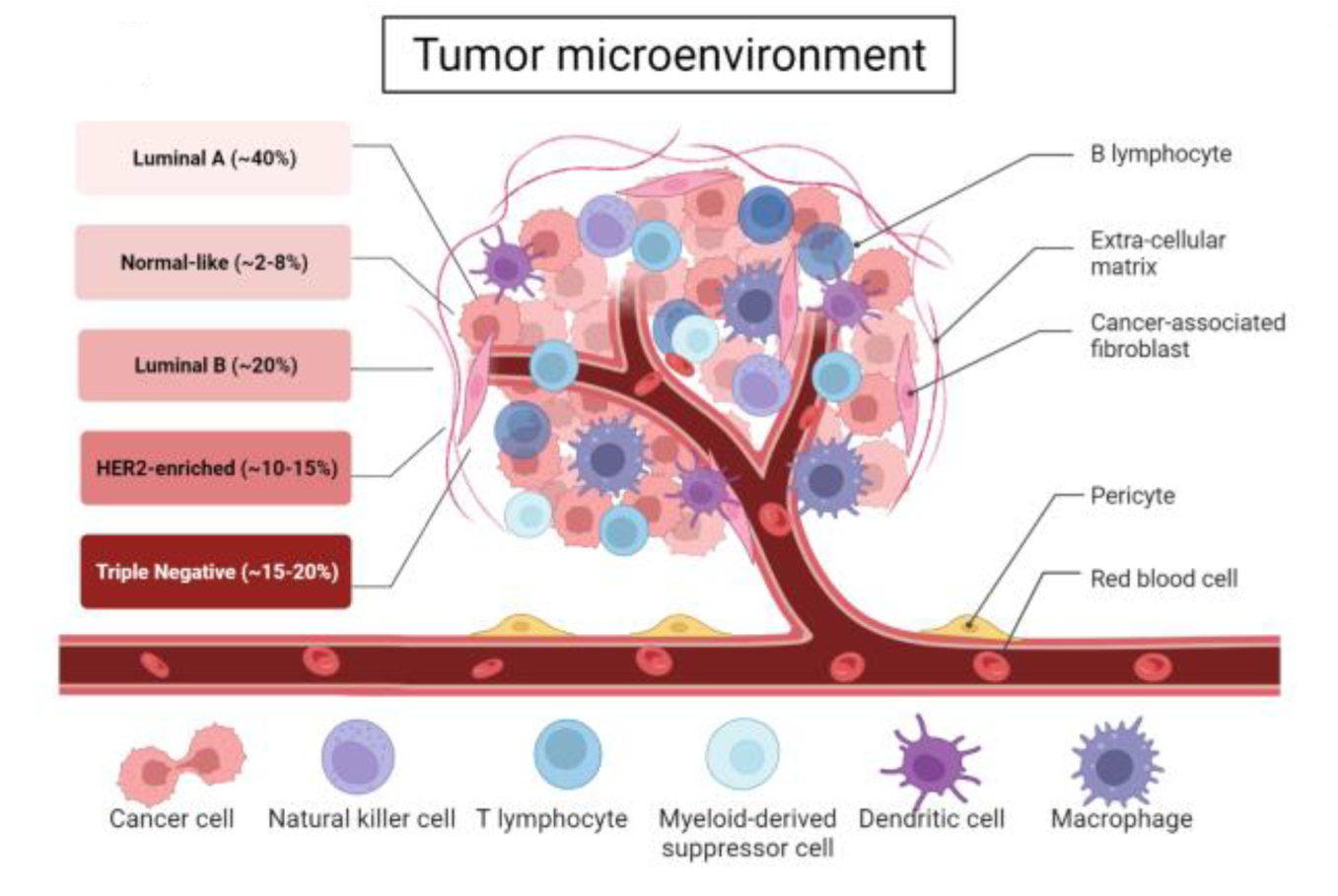
Brožová, K. et al. Proteomes, 2023.
Figure 1. Tumor Microenvironment in Breast Cancer.
Tumor Proteomics Solutions at MtoZ Biolabs
Leveraging high-resolution mass spectrometry-based proteomics platforms, MtoZ Biolabs provides systematic proteomics analysis solutions for tumor samples, enabling the characterization of key molecular features involved in tumor initiation and progression at the protein level.
-
Comprehensive identification of protein composition and proteome profiles in tumor tissues or cell samples
-
Analysis of expression characteristics of specific target proteins or functionally related protein groups
-
Quantitative analysis of tumor proteins to elucidate their expression levels and dynamic changes
-
Comparative analysis of proteomic differences among different sample types, treatment conditions, or experimental groups
This analysis facilitates a systematic understanding of the overall architecture and dynamic alterations of the tumor proteome, providing a reliable foundation for in-depth tumor-related proteomics research.
Workflow of Tumor Proteomics Solutions
1. Sample Processing and Assessment
Quality assessment and basic processing are performed on tumor tissue or cell samples to ensure that sample conditions meet the requirements for proteomics analysis.
2. Protein Extraction and Preprocessing
Total proteins are extracted from samples, followed by interference removal and normalization procedures to improve the consistency and reliability of downstream analyses.
3. Proteolytic Digestion and Mass Spectrometry Analysis
Proteins are subjected to standardized enzymatic digestion, and high-resolution mass spectrometry platforms are used to acquire information on tumor protein composition and expression.
4. Protein Identification and Quantitative Analysis
Based on mass spectrometry data, systematic protein identification and quantitative analysis are conducted to elucidate expression differences across different samples or experimental conditions.
5. Data Integration and Result Output
Analytical results are organized and interpreted to generate structured datasets and analysis reports, providing support for tumor proteomics research.
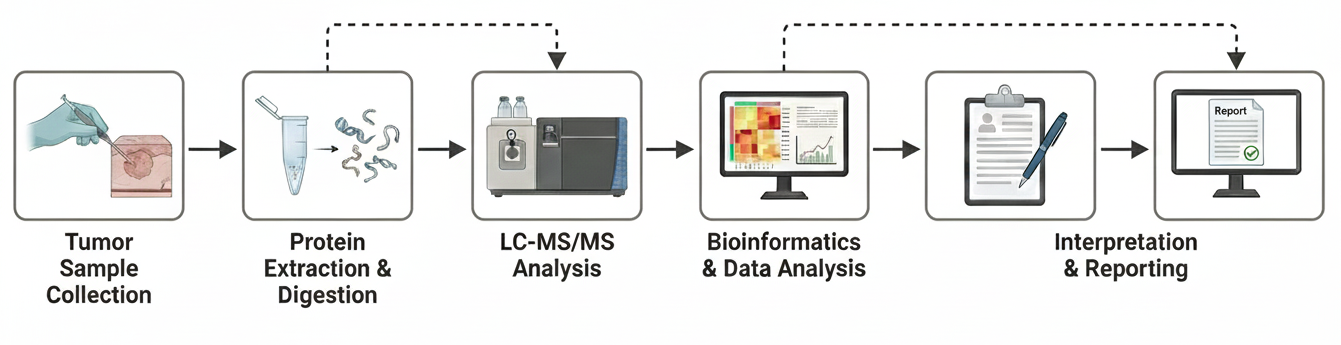
Figure 2. The Tumor Proteimics Analysis Workflow.
Why Choose MtoZ Biolabs?
1. High-Resolution Detection Capability
Facilitates comprehensive characterization of the complex protein composition and expression features in tumor samples.
2. High-Sensitivity Analysis
Suitable for analytical requirements involving strong tumor heterogeneity and a high proportion of low-abundance proteins.
3. Standardized Quality Control System
Strict control of experimental procedures and data processing workflows to ensure result stability and comparability.
4. Customized Analysis Design
Analytical strategies can be flexibly adjusted according to different research stages and objectives.
5. One-Stop Service Support
Covers the entire process from sample assessment to result delivery, improving overall research efficiency and coordination.
Applications of Tumor Proteomics Solutions
1. Tumor Molecular Feature Characterization
Through systematic analysis of protein composition in tumor samples, tumor-associated molecular features and heterogeneity are characterized.
2. Comparative Analysis of Protein Expression
Used for comparative analysis of protein expression patterns across different tumor types, stages, or experimental groups.
3. Signaling Pathway and Regulatory Network Studies
By integrating tumor proteome data, this analysis supports the investigation of key regulatory pathways and functional associations among molecules.
4. Tumor Microenvironment-Related Research
Analyzes protein features in tumor tissues associated with immune components, stromal elements, or cellular interactions, supporting studies of the tumor microenvironment.
5. Drug Development and Mechanism Evaluation
Applied to assess tumor protein changes under drug treatment or experimental interventions, supporting target identification and mechanism-of-action studies.
Deliverables
- Comprehensive Experimental Details
- Materials, Instruments, and Methods
- Tables of protein identification and quantitative analysis results for tumor samples
- Relevant mass spectrometry spectra and visualized analytical figures
- Bioinformatics Analysis
- Raw Data Files
FAQ
Q1: What types of samples are suitable?
A1: Tumor proteomics solutions are primarily applicable to tumor-derived biological samples, including fresh or frozen tumor tissues, tumor samples from different stages or subtypes, as well as properly processed tumor cells or tumor model samples. Samples should maintain good integrity and avoid extensive necrosis, significant protein degradation, or exogenous contamination to ensure the stability, accuracy, and comparability of proteomics analysis results.
Q2: What is the service general workflow?
A2:
Q3: What data formats are provided?
A3: MtoZ Biolabs typically provides standardized deliverables, including:
-
Protein identification and quantitative analysis result tables (Excel/CSV)
-
Related statistical and processed data files
-
Mass spectrometry spectra and visualized analysis results (PNG/TIFF)
-
Raw mass spectrometry data files
-
Comprehensive analysis report (PDF, including an overview of analytical workflows and interpretation of results)
If special analytical requirements exist, data formats can be customized according to project specifications.
Q4: How should I prepare my samples?
A4: To obtain stable and reproducible results, the following points are recommended:
- Sample Quality: Ensure tumor samples are fresh or properly preserved at low temperature, and avoid repeated freeze-thaw cycles and obvious protein degradation.
- Sample Storage: Fresh samples should be frozen as soon as possible; long-term storage is recommended at -80℃.
- Sample Transport: Samples should be transported under low-temperature and sealed conditions to maintain protein stability.
- Additional Information: Providing sample source, tissue type, processing methods, and relevant background information will help optimize the analysis strategy.
For more information, please refer to Sample Submission Guidelines for Proteomics, Sample Submission Guidelines for Metabolomics.
Start Your Project with MtoZ Biolabs
Contact us to discuss your experimental design or request a quote. Whether you are exploring tumor protein expression characteristics and molecular differences or conducting tumor-related mechanistic and functional studies, MtoZ Biolabs can provide reliable analytical support.
