MtoZ Biolabs uses high-resolution mass spectrometry platforms to launch DIA+PRM targeted proteomics services which enables both comprehensive screening of the proteome and precise quantification of specific proteins. This service is widely applied in protein expression change studies, biomarker validation, signaling pathway research, and functional mechanism exploration, providing clients with high-sensitivity, high-specificity quantitative data support.
Overview
DIA (data-independent acquisition) captures all detectable peptide signals from a sample in a single run, providing high-coverage, reproducible global proteomic data; PRM (parallel reaction monitoring) precisely quantifies specific target peptides with high selectivity and sensitivity. The combination of DIA and PRM leverages the advantages of global screening and targeted validation, enabling rapid identification of differential proteins and accurate quantification of key targets, ensuring high reliability of the data.
DIA+PRM Targeted Proteomics Services at MtoZ Biolabs
MtoZ Biolabs provides high-sensitivity, high-selectivity protein quantification services through DIA+PRM targeted proteomics technology.
-
Comprehensive quantification of the proteome
-
Precise quantification of specific target proteins
-
Detection of low-abundance proteins
-
Quantitative comparison between different samples
-
Quantification validation of key targets
Our detection services offer a comprehensive understanding of protein expression changes, enabling researchers to obtain high-precision data support and advance scientific research.
Workflow of DIA+PRM Targeted Proteomics Services
1. Sample Preparation
Protein extraction, enzymatic digestion, and purification are performed to obtain high-quality peptides suitable for mass spectrometry analysis.
2. DIA Data Acquisition
Data-independent acquisition mode is used to comprehensively scan all detectable peptide signals in the sample, providing high-coverage proteomic data.
3. DIA Data Analysis
The DIA data is identified and quantified using a spectral library, selecting target proteins and feature peptides of research value.
4. PRM Targeted Validation
PRM methods are developed for candidate peptides to achieve high specificity and precision in the targeted detection of specific proteins.
5. Integrated Data Analysis
DIA screening results and PRM validation data are combined for comprehensive analysis and visualization, providing reliable quantitative conclusions for research.
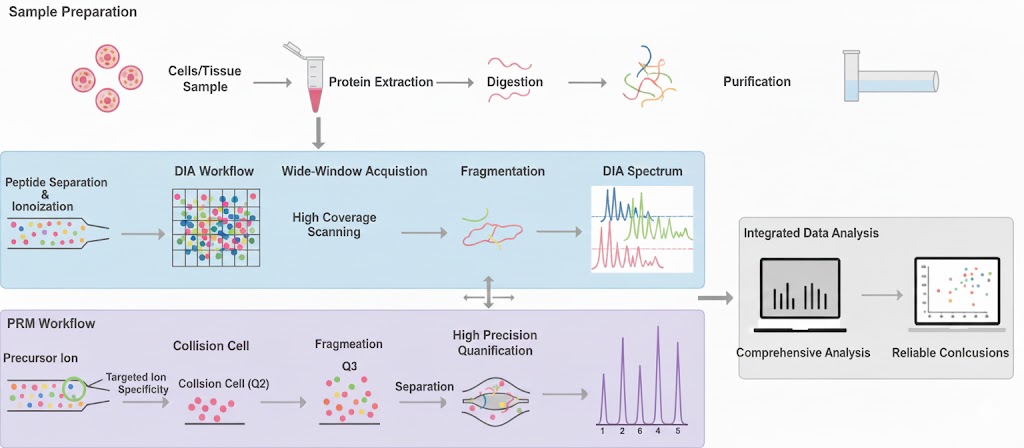
Figure 1. The Workflow of DIA+PRM Targeted Proteomics Analysis.
Why Choose MtoZ Biolabs?
- High Sensitivity Detection: Achieves precise quantification of low-abundance proteins through the combination of DIA and PRM.
- High Selectivity Analysis: PRM provides highly selective detection of specific peptides with low background interference.
- High Quantification Accuracy: Delivers accurate and consistent quantification results even in complex samples.
- Flexible Application: Suitable for various sample types and research needs, supporting large-scale quantitative analysis.
- Rapid Result Output: Optimized analytical workflows ensure the delivery of high-quality data in a short period.
Applications of DIA+PRM Targeted Proteomics Services
1. Biomarker Screening
Quantitative analysis of specific proteins helps identify potential biomarkers related to diseases or biological processes.
2. Drug Target Research
Targeted protein quantification supports the screening, validation, and efficacy evaluation of drug targets.
3. Environmental Stress Research
Suitable for studying protein changes under environmental shifts or stress conditions, helping understand biological responses to external environments.
4. Cell Signaling Pathway Analysis
Analyzing the expression changes of key proteins in signaling pathways, aiding in understanding cellular regulatory mechanisms.
5. Post-Translational Modification Analysis
Used to study protein post-translational modifications, such as phosphorylation and acetylation, helping reveal functional regulatory mechanisms.
Deliverables
- Comprehensive Experimental Details
- Materials, Instruments, and Methods
- Protein Quantification Results
- Visualization Files
- Bioinformatics Analysis
- Raw Data Files
FAQ
Q1: What types of samples are suitable?
A1: DIA+PRM targeted proteomics services are suitable for proteomic samples with a clear origin, high purity, and good solubility, including cell lysates, tissue extracts, serum, plasma, and recombinant proteins. For low-abundance or complex matrix samples, preprocessing is recommended to improve the accuracy of targeted detection.
Q2: What is the service general workflow?
A2:
Q3: What data formats are provided?
A3: MtoZ Biolabs provides multiple standardized data output formats, including:
-
Raw mass spectrometry data files (e.g., .d/.raw platform formats)
-
Quantification results for target proteins (Excel/CSV formats)
-
Analysis report PDF (including experimental methods, data summary, and result interpretation)
-
Visualization files (e.g., chromatogram peak diagrams, fragment ion spectra, PNG/TIFF formats)
If special analytical requirements exist, data formats can be customized according to project specifications.
Q4: How should I prepare my samples?
A4: To ensure optimal quantification results, we recommend:
- Sample Purity: Avoid components such as high salt, detergents, and glycerol that may interfere with analysis.
- Sample Storage: Store samples at 4℃ for short-term use, and at -80℃ for long-term storage, avoiding repeated freeze-thaw cycles.
- Sample Transport: Use dry ice or ice packs for cold transport and ensure sealed packaging to prevent leakage or contamination.
- Additional Information: Please provide sample type, concentration, buffer system, and pretreatment steps, so we can tailor the most appropriate analysis strategy to your needs.
For more information, please refer to Sample Submission Guidelines for Proteomics, Sample Submission Guidelines for Metabolomics.
Start Your Project with MtoZ Biolabs
Contact us to discuss your experimental design or request a quote. Whether you are exploring changes in the expression of target proteins or studying quantitative differences in complex systems, we can provide you with high-quality, reproducible proteomics analysis support.
