MtoZ Biolabs provides comprehensive Plant Proteomics Analysis Service to support global protein identification, quantitative comparison, protein interaction studies, and post-translational modification-related research in plant systems. Built on standardized protein extraction procedures and high-resolution mass spectrometry platforms, this service enables flexible experimental design aligned with specific research objectives.
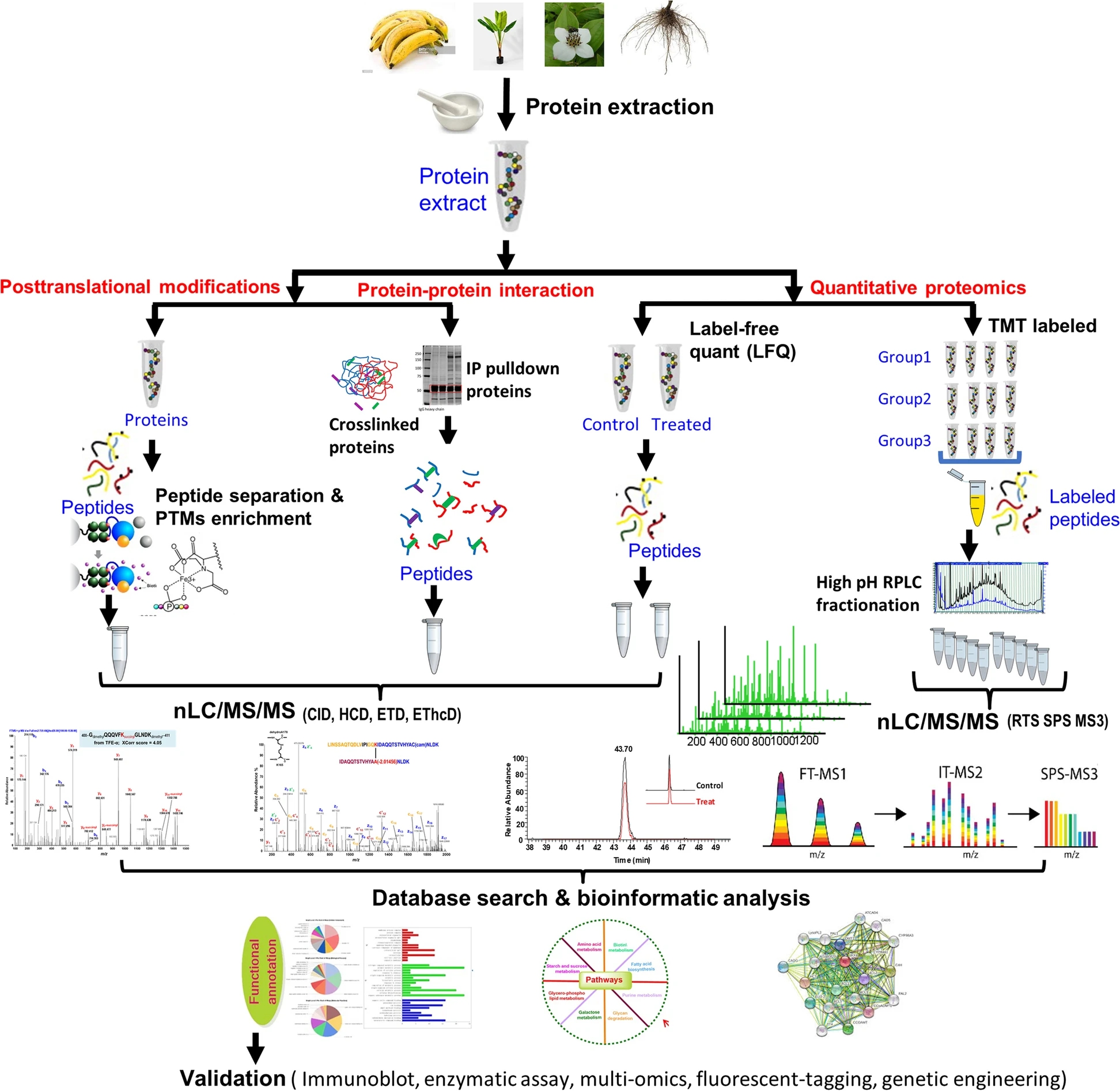
Yan, S. et al. Mol Hortic. 2022.
Figure 1. Plant Proteomics Analysis Workflows
Overview
Plant proteomics focuses on the systematic analysis of protein composition and abundance in plant tissues under defined biological conditions. Starting from reliable protein extraction, proteomic analysis can be configured to address different experimental questions, including expression profiling, quantitative comparison between groups, interaction analysis, and modification-focused studies.
By selecting appropriate sample preparation routes and mass spectrometry acquisition strategies, plant proteomics provides protein-level evidence to support functional studies, mechanism exploration, and phenotype-associated research in plant biology.
Plant Proteomics Analysis Service at MtoZ Biolabs
MtoZ Biolabs offers an integrated plant proteomics platform in which analytical strategies can be selected and combined based on project needs.
1. Global Plant Proteome Profiling
Large-scale identification of proteins expressed in plant tissues, generating comprehensive proteomic datasets for baseline or condition-specific studies.
2. Quantitative Plant Proteomics
Comparative analysis of protein abundance across experimental groups using label-free or isobaric labeling strategies, supporting differential expression analysis.
3. Plant Post-Translational Modifications Proteomics
Proteomics workflows designed to accommodate downstream or parallel analysis of modified peptides, enabling investigation of regulatory protein modifications.
4. Protein-Protein Interaction Proteomics
Identification of interacting proteins or protein complexes following affinity enrichment or crosslinking-based experimental designs.
5. Targeted Protein Analysis within Complex Samples
Focused evaluation of selected proteins of interest within global proteomic datasets to support hypothesis-driven studies.
Why Choose MtoZ Biolabs?
- High-Resolution Mass Spectrometry Platforms: Advanced LC-MS/MS systems supporting discovery-driven and quantitative plant proteomics.
- Experience with Plant Sample Complexity: Proven handling of plant tissues rich in polysaccharides, phenolics, and secondary metabolites.
- Quantitative Strategy Options: Support for label-free and multiplexed quantitative designs based on experimental scale.
- Flexible Proteomics Configuration: Analytical routes can be adjusted according to biological questions rather than restricted to a single workflow.
- One-Time-Charge: Our pricing is transparent, no hidden fees or additional costs.
Start Your Project with MtoZ Biolabs
MtoZ Biolabs delivers reliable and scalable Plant Proteomics Analysis Service to support functional plant biology research. Contact us to discuss your study design or request a customized proteomics solution tailored to your research goals.
FAQ
Q1: What types of samples are suitable?
- Fresh Tissues: Leaves, roots, stems, flowers, seeds, etc.; at least 100 mg
- Dried Tissues: Plant tissues processed by freeze-drying or drying; at least 50 mg
- Cells: Plant cell suspension or cell lysate; at least 1x107 cells
- Protein Extracts: Protein samples extracted, at least 100 μg with their concentration not less than 1 μg/μL
For more information, please refer to Sample Submission Guidelines for Proteomics. If you are unsure about sample preparation, our team is available for consultation and can guide you through the process to ensure optimal results.
Q2: What is the service general workflow?
Q3: What data formats are provided?
Deliverables include raw LC-MS/MS data files, processed protein identification and quantification tables in Excel or CSV format, and a comprehensive PDF report summarizing analytical methods, quality control, and key findings. Figures and visualizations are provided in PNG or TIFF format.
Customized data formats or publication-ready outputs can be provided upon request to meet specific research or journal requirements.
Related Services
