MtoZ Biolabs offers Competitive Activity-Based Protein Profiling (ABPP) Service to enable functional interrogation of enzyme activities and small-molecule target engagement in complex biological systems. This service is designed to support mechanism-of-action studies, target identification, and selectivity assessment for bioactive compounds at the proteome-wide level.
What Is Competitive Activity-Based Protein Profiling?
Competitive activity-based protein profiling is a functional proteomics strategy that measures enzyme activity rather than protein abundance. In a competitive ABPP experiment, a bioactive compound of interest is introduced prior to labeling with an activity-based probe. If the compound engages an enzyme active site, it prevents probe binding, resulting in a measurable reduction of probe labeling.
By directly linking chemical inhibition to enzymatic activity, competitive ABPP enables the identification of functional targets, off-target interactions, and pathway-level effects in native biological contexts. This approach is particularly valuable for profiling enzyme families with conserved structures, where affinity-based methods may lack specificity.
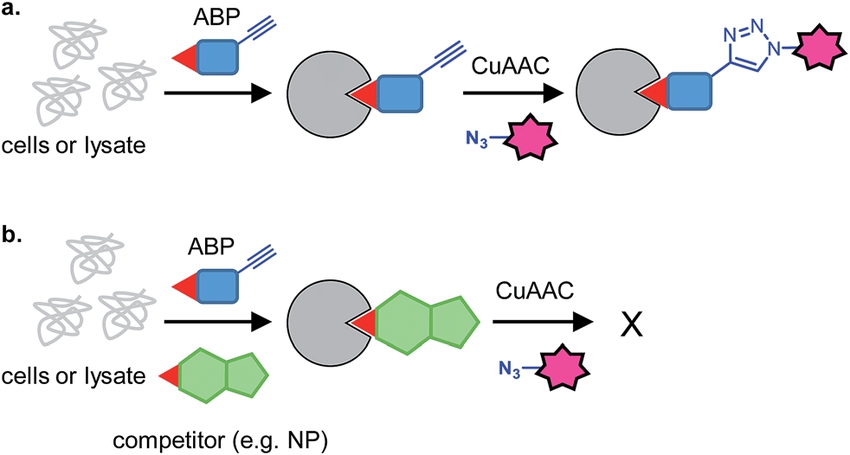
Wright, M. H. et al. Nat. Prod. Rep. 2016.
Figure 1. Principle of Competitive ABPP
Competitive Activity-Based Protein Profiling (ABPP) Service at MtoZ Biolabs
MtoZ Biolabs provides Competitive ABPP workflows that can be tailored to different enzyme classes and experimental objectives.
1. Target Engagement Assessment
Evaluation of whether a test compound directly engages its intended protein targets in complex proteomes.
2. Inhibitor Selectivity Profiling
Comparative analysis of probe labeling patterns to identify on-target and off-target interactions across enzyme families.
3. Activity-Based Target Identification
Discovery of functionally active proteins affected by competing ligands, supporting mechanism-of-action studies.
4. Comparative Profiling Across Conditions
Assessment of compound-protein interactions across different treatment groups, concentrations, or biological states.
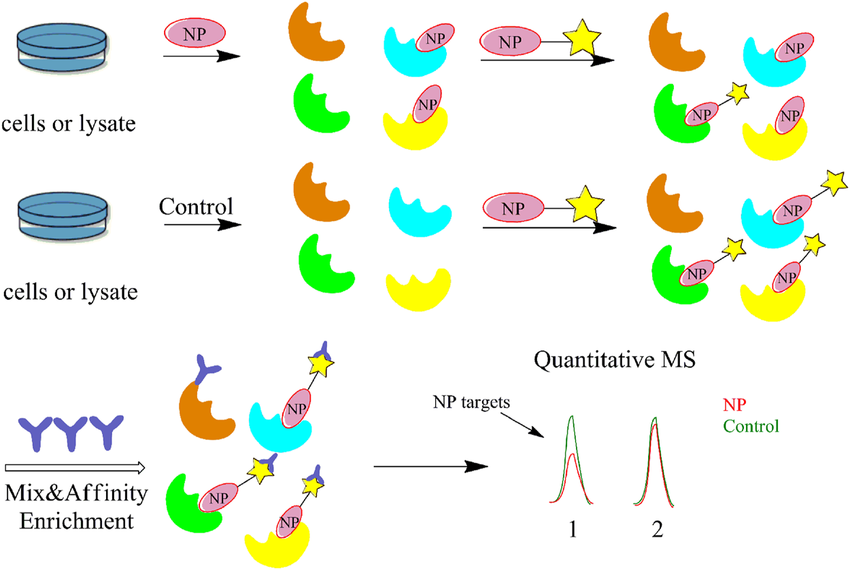
Li, G. et al. Front. Chem. 2021.
Figure 2. Flow Diagram of the Competitive Activity-Based Protein Profiling (ABPP) Strategy
Why Choose MtoZ Biolabs?
✔ Support for Drug Discovery Studies
Well-suited for target validation, lead optimization, and selectivity assessment.
✔ Flexible Experimental Design
Workflows can be adapted to different probe chemistries, enzyme classes, and study goals.
✔ Compatibility with Complex Proteomes
Applicable to cell lysates, tissue samples, and other native biological matrices.
✔ Clear and Interpretable Deliverables
Results highlight functionally engaged targets and potential off-target interactions.
✔ One-Time-Charge
Our pricing is transparent, no hidden fees or additional costs.
Applications of Competitive Activity-Based Protein Profiling (ABPP) Service
1. Target Engagement Verification
Confirm whether a small molecule truly binds its intended protein target in native biological systems.
2. Off-Target and Selectivity Profiling
Identify unintended protein interactions that may contribute to toxicity or side effects.
3. Mechanism-of-Action Studies
Elucidate how compounds modulate enzymatic activity within signaling or metabolic pathways.
4. Lead Optimization Support
Compare binding selectivity and potency across compound series during medicinal chemistry campaigns.
Start Your Project with MtoZ Biolabs
MtoZ Biolabs provides Competitive Activity-Based Protein Profiling services to support functional proteomics and small-molecule research. Whether your goal is target validation, selectivity assessment, or mechanism exploration, our Competitive ABPP Service delivers activity-centered insights to guide decision-making. Contact us to discuss your project design and experimental requirements.
FAQ
Q1: What types of samples are suitable?
Competitive ABPP can be performed on cell lysates, tissue extracts, or other complex protein samples. Sample compatibility depends on the enzyme class and probe chemistry and is evaluated during project consultation.
Q2: How should I prepare my samples?
Samples should be prepared under conditions that preserve native protein activity and minimize background interference. General considerations include:
-
Avoiding harsh denaturing agents that may compromise enzymatic activity.
-
Controlling temperature, handling time, and storage conditions.
-
Using buffers compatible with activity-based probe labeling.
For more information, please refer to Sample Submission Guidelines for Proteomics. If you are unsure about sample preparation, our team is available for consultation and can guide you through the process to ensure optimal results.
Q3: What is the service general workflow?