MtoZ Biolabs provides Tear Proteomics Solutions designed for deep protein profiling from tears. Tear fluid contains secreted defense proteins, enzymes, structural epithelial proteins, immune mediators, and trace systemic components, enabling high-value molecular reads from the corneal and conjunctival surface. MtoZ Biolabs converts small clinical volumes into confident protein identities and reproducible quantitative matrices for biomarker discovery, ocular host response programs, and mechanism-focused clinical proteomics research.
Background
Tear fluid is a non-invasive, clinically valuable biofluid containing secreted proteins, enzymes, immune mediators, structural ocular-surface proteins, and trace plasma-derived components. These proteins form the biochemical foundation of corneal protection, lubrication, epithelial homeostasis, antimicrobial defense, and inflammatory signaling. The tear proteome rapidly adjusts to infection, allergy, dryness, systemic metabolic imbalance, and ocular surface disruption, making it highly suitable for linking molecular protein shifts to clinical phenotypes.
Tear-proteomics has become a powerful route for identifying biomarkers associated with conditions such as dry eye disease, keratoconjunctivitis, blepharitis, allergic eye inflammation, viral or bacterial conjunctival infections, corneal injury, systemic autoimmune-related ocular involvement, and diabetic ocular-surface stress. The protein landscape within tears supports both early molecular stratification and longitudinal monitoring of disease-associated expression programs.
Tear Proteomics Solutions at MtoZ Biolabs
MtoZ Biolabs delivers Tear Proteomics Solutions through a unified mass spectrometry pipeline that supports DIA for deep, unbiased protein detection, targeted panels for precise quantification, and discovery proteomics for broad ocular proteome coverage from tear matrices.
The service reveals inflammatory mediators, antimicrobial proteins, epithelial defense proteins, enzymes, and regulatory nodes suitable for biomarker screening and ocular mechanism research. You only need to submit your samples, and we manage every stage including extraction, digestion, LC-MS acquisition, data processing, and interpretation.
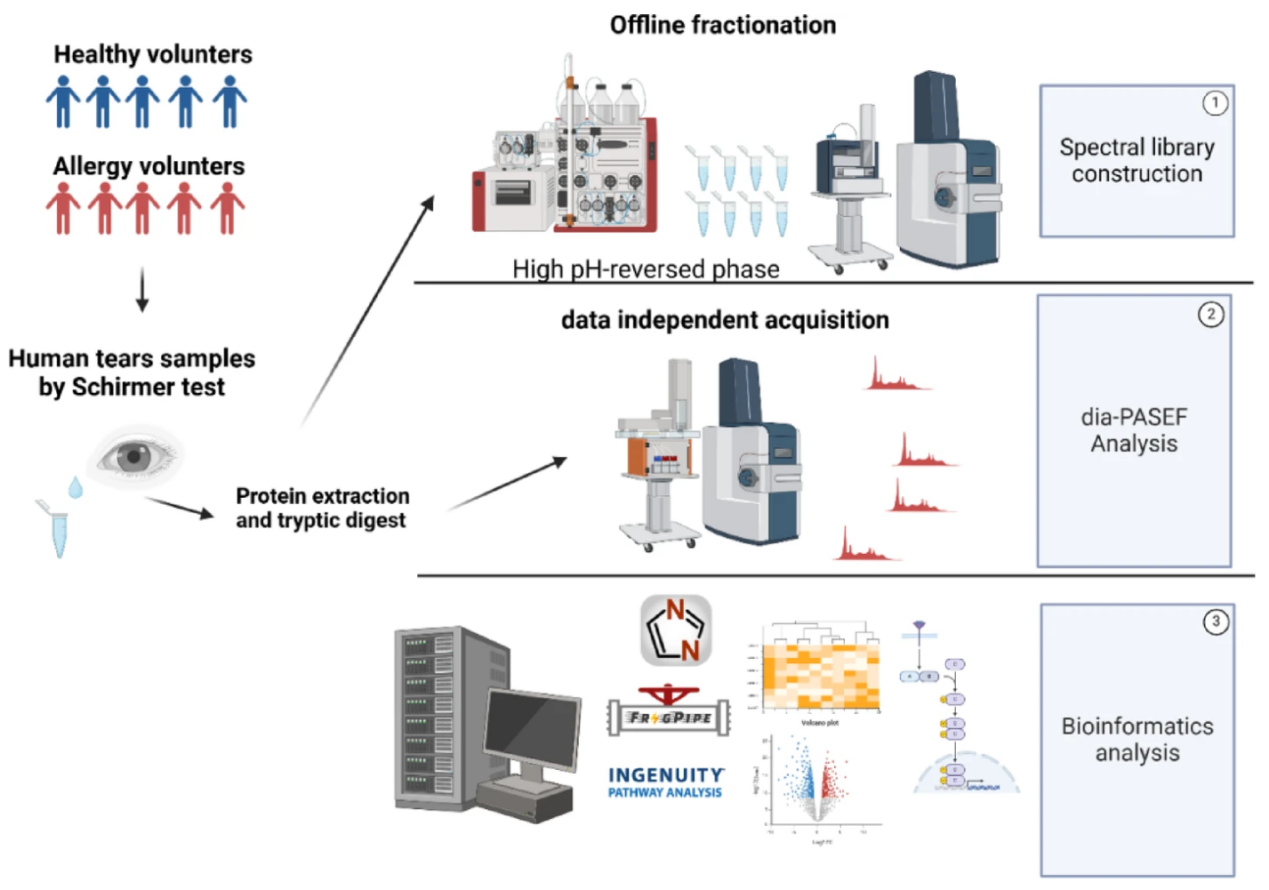
Vera-Montecinos, A. et al. Sci Rep. 2025.
Figure 1. Experimental Design to Study the Proteomic Profile of the Human Tear's Samples
Why Choose MtoZ Biolabs
- High-resolution MS platforms and calibrated acquisition ensure accurate protein identification and quantification.
- DIA-enabled runs deliver fast, scalable processing for large tear-proteome studies.
- Established expertise in proteomics brings strong confidence even from micro-volume clinical samples.
- Targeted panels and discovery depth are customized to match specific research objectives.
- Responsive coordination and clear guidance support smooth project execution and delivery.
Applications of Tear Proteomics Solutions
1. Ocular Surface Disease Biomarker Discovery
Tear proteomics identifies protein signatures associated with dry eye, infection, allergy, or corneal surface disruption, supporting early molecular detection and progression monitoring.
2. Infection-Triggered Host Protein Response
Protein shifts in tear matrices during viral or bacterial infections reveal mucosal defense behavior and host immune protein programs for mechanism-level studies.
3. Environmental Ocular Stress Proteome Adjustments
Protein alterations triggered by airborne or occupational irritants, allergens, or pollutants reveal tear proteome defensive adjustments for exposure-linked proteomics research.
FAQ
Q1: What types of samples are suitable?
Purified protein, cultured cells, and frozen or freshly collected tear fluid are all compatible.
Q2: How should I prepare my samples?
- Collect tears using capillary tubes, tear strips, or swab elution and transfer them into low-binding tubes.
- Snap-freeze samples immediately after collection to preserve low-abundance tear proteins and prevent degradation.
- Avoid detergents, high salts, strong buffers, or reactive chemicals that may interfere with LC-MS/MS analysis.
- Store samples at −80 °C and ship on dry ice to maintain protein stability throughout transport.
Detailed requirements and recommendations can be found in the Sample Submission Guidelines for Proteomics.
Q3: What is the service general workflow?
Q4: What data formats are provided?
- Raw mass spectrometry files such as .raw, .d, or .wiff.
- Quantitative result tables in .xlsx or .csv, including intensity and key statistical or database-derived scores.
- Bioinformatics summaries exported in .xlsx or .csv for pathway and interaction views.
- Publication-ready figures in high-resolution .png or .tiff.
- A complete PDF report summarizing methods and QC metrics.
- Extra formats are available on request.
Start Your Project with MtoZ Biolabs
MtoZ Biolabs reveals the tear proteome with sensitivity, rigor, and reproducible depth, turning small clinical tear volumes into reliable protein identities and quantitative readouts for clinical research. Connect with MtoZ Biolabs for free business assessment to start your tear proteomics study.
