MtoZ Biolabs provides SILAC-based Quantitative Proteomics Analysis Service using metabolic stable isotope labeling combined with high-resolution LC-MS/MS to deliver accurate relative protein quantification in cell systems. This service supports mechanism studies, signaling analysis, and biomarker discovery where precise comparison of proteomes between defined cellular states is required.
What Is SILAC?
SILAC (Stable Isotope Labeling by Amino acids in Cell culture) is a metabolic labeling strategy for quantitative proteomics. Cells are grown in media containing either natural "light" amino acids or "heavy" isotopically labeled amino acids such as 13C- or 15N-labeled lysine and arginine. Over several cell doublings, the labeled amino acids are incorporated into newly synthesized proteins, resulting in complete or near-complete labeling of the cellular proteome.
In a typical SILAC experiment:
- One condition (for example control) is cultured in light medium.
- Another condition (for example treated or mutant) is cultured in heavy medium.
- Cells are harvested, proteins are extracted and mixed at a 1:1 ratio between light and heavy conditions.
- The combined protein mixture is then analyzed by LC-MS/MS.
Because light and heavy peptides are chemically identical but differ in mass, they co-elute in chromatography but appear as distinct peaks in the mass spectrum. The intensity ratios of heavy and light peptide pairs provide a direct and accurate measure of relative protein abundance between conditions.
Because labeling occurs at the cellular level, SILAC minimizes variation from sample handling, digestion, or LC-MS, making it one of the most accurate strategies for relative quantification in cultured cells.
SILAC-Based Quantitative Proteomics Analysis Service at MtoZ Biolabs
MtoZ Biolabs offers SILAC-based Quantitative Proteomics Analysis Service built on high-resolution Orbitrap platforms such as Q Exactive HF and Orbitrap Fusion Lumos combined with nano-LC and optimized SILAC workflows. The service is designed for cell models where metabolic labeling is feasible and high-precision quantification is required.
Our SILAC-based solution supports:
- Two-state and multi-state comparisons (for example control vs treated, wild-type vs mutant).
- Time course studies using pulse SILAC or multi-label designs.
- Mechanistic studies of signaling, proteome remodeling, and protein complex dynamics.
You only need to provide your samples and project objectives. MtoZ Biolabs handles the entire SILAC workflow, including cell culture labeling with stable isotopes, protein extraction, enzymatic digestion, LC-MS/MS analysis, quantitative data processing, and downstream bioinformatics interpretation, delivering ready-to-use results that directly address your biological questions.
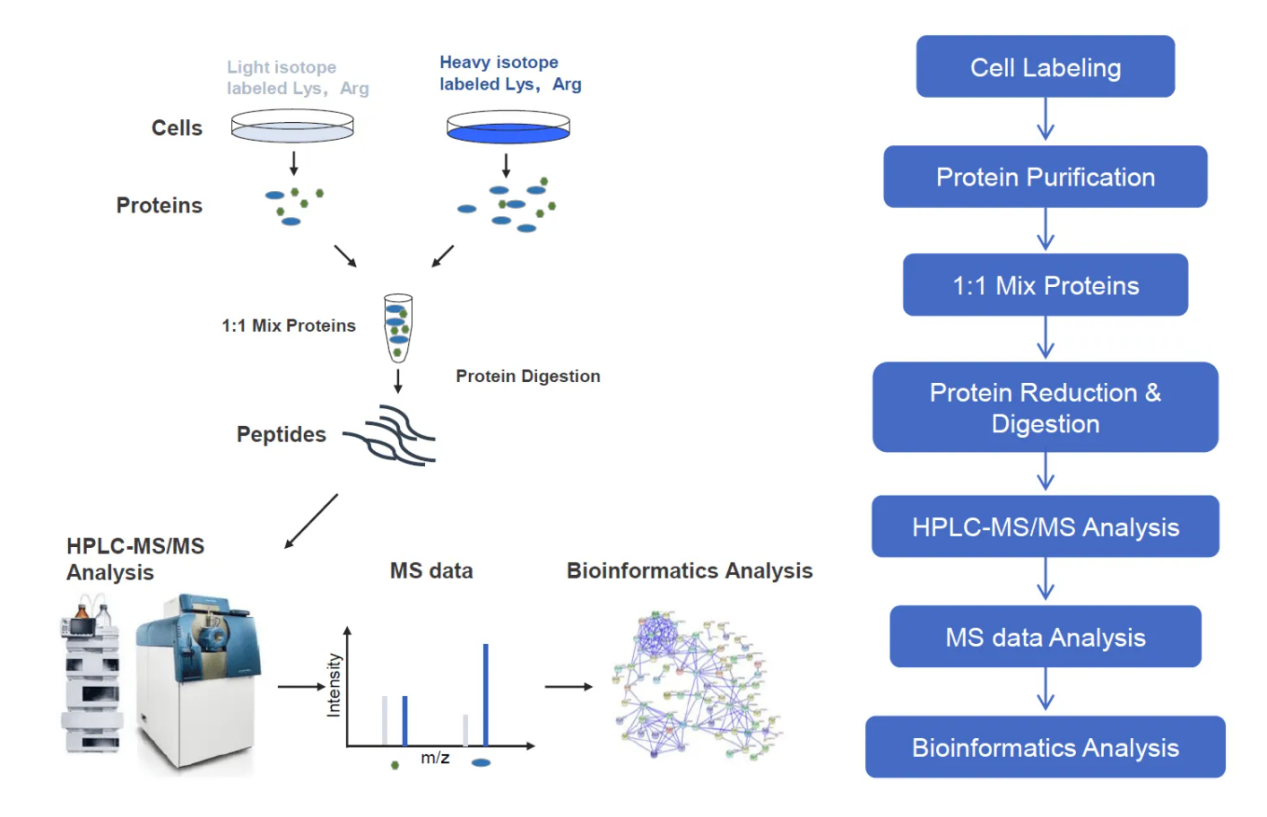
Figure 1. Workflow for SILAC-based Quantitative Proteomics Analysis Service
Why Choose MtoZ Biolabs
1. Flexible Experimental Support
Assistance with label scheme design, time points, and replicate structure tailored to your model system.
2. End-to-End Service
From study planning and sample processing to final figures and summary reports, a dedicated proteomics team supports every stage of your project.
3. High-Precision Quantification
Metabolic labeling combined with high-resolution Orbitrap MS supports accurate heavy/light ratio measurements and low technical variation.
4. Optimized SILAC Workflows
Established protocols for cell lysis, digestion, and LC-MS ensure reproducible results across replicates and projects.
5. Deep Proteome Coverage
Nano-LC and advanced acquisition methods enable quantification of thousands of proteins in a single experiment.
Applications of SILAC-Based Quantitative Proteomics Analysis Service
SILAC-based Quantitative Proteomics Analysis Service at MtoZ Biolabs can be applied to many cell-based research scenarios.
1. Signal Transduction and Pathway Analysis
Quantify changes in kinases, phosphatases, and signaling effectors after stimulation or inhibition.
2. Drug Mechanism and Target Validation
Compare proteomes of treated vs untreated cells to uncover direct and indirect drug effects.
3. Protein Complex and Interaction Dynamics
Combine SILAC labeling with affinity purification to study changes in complex composition.
4. Cell Cycle, Differentiation, and Stress Response
Characterize proteome shifts across cell cycle phases, differentiation stages, or stress conditions.
FAQ
Q1: What types of samples are suitable?
- Viable adherent or suspension cell lines suitable for culture and expansion
- Stable engineered cell lines (for example overexpression or knockdown models)
- Primary or stem cell lines that can be adapted to SILAC medium after feasibility assessment
SILAC is designed for cell culture systems and is generally not applied directly to primary tissues or biofluids.
Q2: How should I prepare my samples?
- Send frozen cell vials or actively growing cultures with clear documentation of medium and culture conditions
- Ensure cells are mycoplasma free and in good viability before shipment
For more information, please refer to Sample Submission Guidelines for Proteomics.
Q3: What is the service general workflow?
Q4: What data formats are provided?
- Raw LC-MS/MS files in instrument specific formats
- Peptide and protein identification tables with SILAC heavy/light ratios in .xlsx or .csv
- Normalized protein abundance matrices and differential expression results in .xlsx or .csv
- Functional and pathway analysis outputs in spreadsheet format
- Publication ready figures such as volcano plots, heatmaps, and PCA plots in high-resolution PNG or TIFF
- A concise PDF report summarizing experimental design, SILAC setup, methods, QC metrics, and key quantitative findings
- Additional or custom formats can be provided on request
Start Your Project with MtoZ Biolabs
Our SILAC-based Quantitative Proteomics Analysis Service is designed to provide more rapid, high-throughput, and cost-effective analysis, with exceptional data quality and minimal sample consumption. Free project evaluation, welcome to learn more details.
