MtoZ Biolabs provides Label-free Quantitative Proteomics Analysis Service using high-resolution LC-MS/MS to compare protein expression across biological conditions without the need for chemical or isotope labels. This service delivers large scale protein identification, accurate relative quantification, and functional interpretation for discovery research, biomarker screening, and mechanism studies.
What Is Label-free Quantitative Proteomics?
Label-free quantitative proteomics is a mass spectrometry based strategy that measures protein abundance directly from MS signals instead of using isotopic or isobaric labels. It is widely used to compare proteomes between different samples, time points, treatments, or phenotypes in a flexible and scalable way.
Key features of label-free quantitative proteomics include:
1. No Labeling Step Required
-
Samples are processed individually without metabolic or chemical labeling
-
Suitable for primary tissues, clinical samples, and precious or limited materials
2. Flexible Experimental Design
-
Easy to increase or decrease the number of samples or groups
-
Ideal for time course studies, large cohorts, and pilot screens
3. Global Proteome Coverage
-
Enables simultaneous identification and quantification of hundreds to thousands of proteins
-
Captures broad changes in signaling, metabolism, structure, and regulation
Principle of Label-free Quantitative Proteomics
Label free quantitative proteomics infers protein abundance directly from mass spectrometry signals without introducing isotope or chemical tags. By extracting quantitative information from peptide ions across LC-MS runs, it enables robust comparison of protein expression between experimental groups. Two strategies are commonly used in this workflow.
1. Intensity Based Quantification
In intensity based label free analysis, peptide abundance is estimated from the ion signal intensity or integrated peak area in the mass spectrum. After alignment of retention time, peak picking, and normalization across runs, these values are aggregated at the protein level to generate quantitative profiles. This strategy is well suited for complex samples, offering broad dynamic range, high sensitivity, and the ability to track subtle changes and low abundance proteins across multiple conditions.
2. Spectral Counting
In spectral counting approaches, protein abundance is approximated using the number of MS/MS spectra assigned to peptides from a given protein. Proteins that produce more identified spectra are interpreted as more abundant. When combined with appropriate normalization and statistical models, spectral counting provides an efficient and straightforward way to compare protein levels, particularly for large-scale proteomic studies and high-abundance protein quantification.
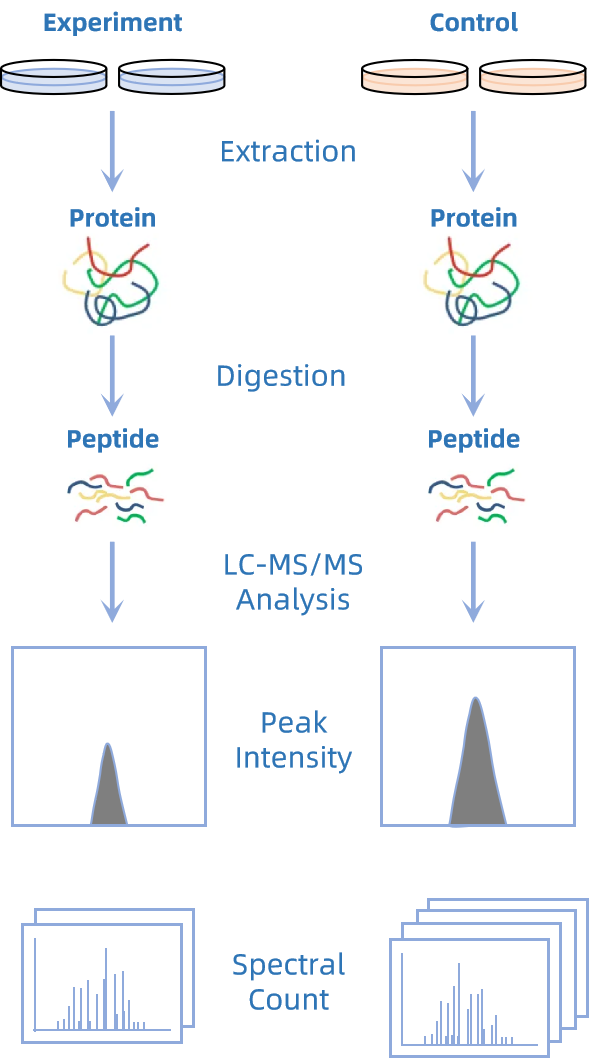
Figure 1. Principle of Label-free Quantitative Proteomics Analysis
Label-free Quantitative Proteomics Analysis Service at MtoZ Biolabs
MtoZ Biolabs offers Label-free Quantitative Proteomics Analysis Service built on high resolution LC-MS/MS platforms and optimized sample preparation workflows. Our service is designed to generate robust, comparable protein profiles across multiple groups and biological replicates. You only need to submit your samples, and MtoZ Biolabs manages extraction, digestion, LC-MS acquisition, data processing, and biological interpretation.
Workflow of Label-free Quantitative Proteomics Analysis Service
- Sample Preparation: Extract proteins from cells, tissues, or biofluids using compatible lysis and cleanup.
- Enzymatic Digestion: Reduce, alkylate, and digest equal protein amounts with sequencing grade proteases to generate peptides.
- LC-MS/MS Analysis: Separate peptides by LC and acquire high-resolution MS/MS data using label free DDA or DIA methods.
- Data Analysis: Identify proteins, perform label free quantification, run statistical comparisons, and summarize functional insights.
Why Choose MtoZ Biolabs
- Flexible experimental design supports small pilot sets through large multi group or time course studies without label constraints.
- Robust QC and normalization strategies control run to run variation, improving confidence in differential protein expression results.
- Broad proteome coverage enables simultaneous quantification of thousands of proteins, including low abundance regulatory and signaling molecules.
- Experienced proteomics scientists assist with study design, sample strategy, and data interpretation tailored to your biological questions.
Applications of Label-free Quantitative Proteomics Analysis Service
Label-free quantitative proteomics can be applied in many research areas:
1. Biomarker Discovery
-
Identify proteins that differ between disease and control groups
-
Generate candidate biomarkers for follow up validation
2. Mechanism and Pathway Studies
-
Characterize proteomic changes following drug treatment, knockdown, overexpression, or environmental stress
-
Map proteins into pathways to infer mechanisms of action
3. Comparative Proteomics Across Models
-
Compare proteomes across cell lines, tissues, organoids, or animal models
-
Study species differences and translational relevance
4. Time Course and Dynamic Response
-
Monitor protein expression over time during development, differentiation, or treatment
-
Track early and late protein changes in dynamic processes
5. Support for Multi Omics Projects
-
Integrate label-free proteomics with genomics, transcriptomics, or metabolomics
-
Build more complete views of biological systems and disease states
FAQ
Q1: What types of samples are suitable?
We support a wide range of label free proteomics samples, including:
-
Cell pellets from adherent or suspension cell lines
-
Primary cells from blood, tissues, or organoids
-
Fresh frozen tissues or biopsies
-
Biofluids such as serum, plasma, CSF, urine, and others
-
Microbial cultures or model organism samples
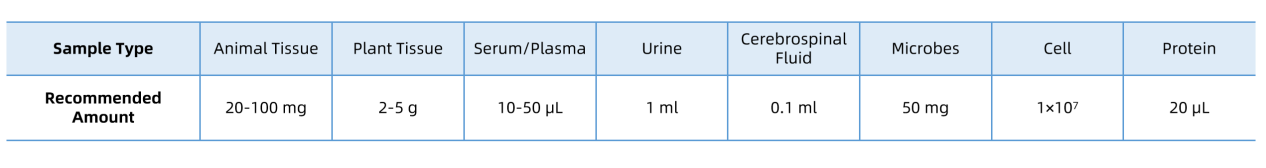
For unusual sample types or low-yield materials, please contact MtoZ Biolabs for customized preparation guidance.
Q2: How should I prepare my samples?
- Use clean, low binding tubes and avoid repeated freeze thaw cycles
- Snap freeze samples after collection and store at −80 °C
- Avoid strong detergents, very high salt, and large amounts of reducing agents
- Record detailed metadata (sample type, group, treatment, time point)
- Ship on dry ice with a clearly labeled sample list
Detailed requirements and recommendations can be found in the Sample Submission Guidelines for Proteomics.
Q3: What is the service general workflow?
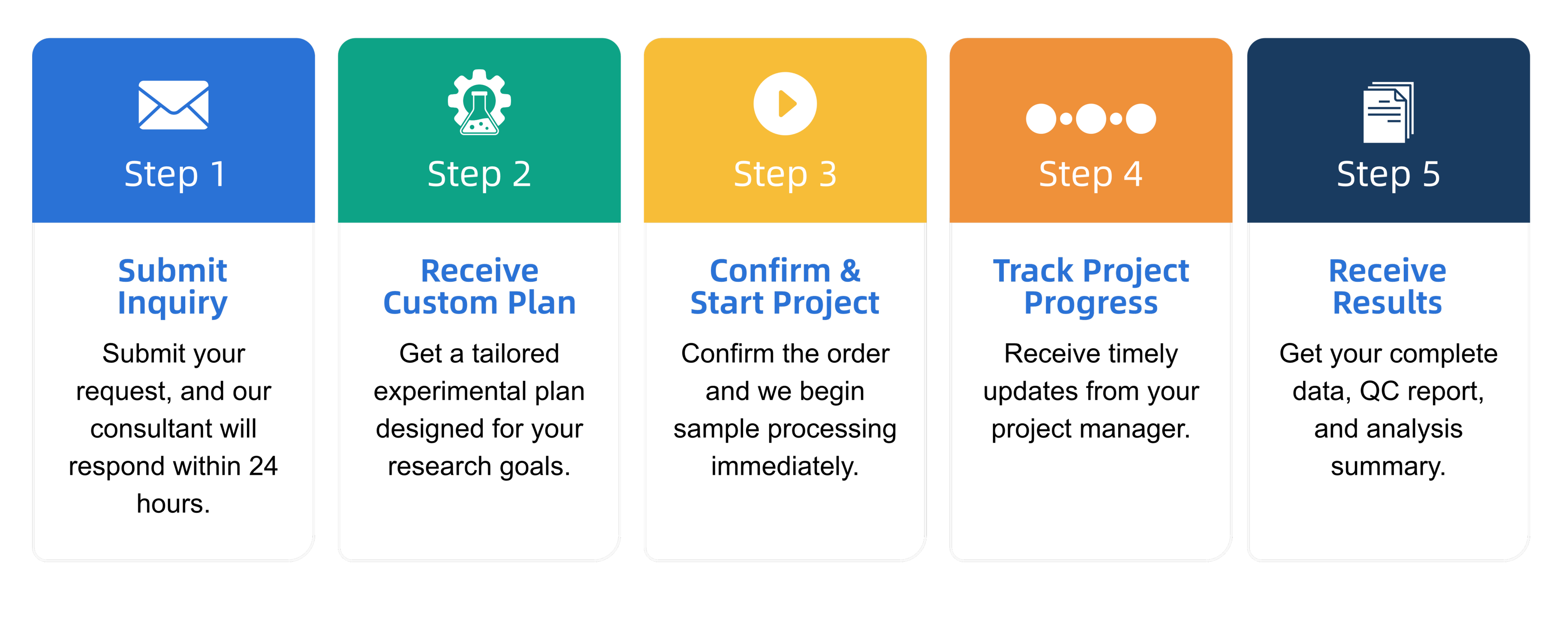
Q4: What data formats are provided?
- Raw LC-MS/MS files in instrument specific formats
- Peptide and protein identification and quantification tables in .xlsx or .csv
- Statistical and functional analysis outputs in .xlsx or .csv
- Publication ready figures in high resolution .png or .tiff
- A concise PDF report summarizing methods, QC metrics, and key findings
- Additional formats can be provided on request
Start Your Project with MtoZ Biolabs
MtoZ Biolabs Label-free Quantitative Proteomics Analysis Service provides a powerful solution for unbiased protein expression profiling without labeling constraints.
Contact MtoZ Biolabs for a free business assessment.
