MtoZ Biolabs offers comprehensive proteomics solutions designed to advance cancer research by uncovering protein-level changes associated with tumor biology, progression, and treatment response. Our advanced mass spectrometry-based workflows provide detailed insights into the molecular mechanisms driving cancer, enabling the identification of potential biomarkers and therapeutic targets with high sensitivity and reproducibility.
Introduction
Cancer is driven by alterations in protein expression, post-translational modifications, signaling networks, and microenvironmental interactions that cannot be fully captured at the genomic or transcriptomic level alone. Proteomics provides a functional, phenotype-proximal view of these biological processes, enabling researchers to analyze dynamic protein landscapes, identify disease-specific molecular signatures, and uncover mechanistic vulnerabilities for therapeutic intervention.
Key contributions of cancer proteomics include:
1. Understanding Disease Mechanisms
Proteomic profiling reveals:
-
Dysregulated pathways linked to proliferation, apoptosis, metabolism, DNA repair, angiogenesis, and metastasis
-
Activation states of kinases and signaling modules through phosphorylation, acetylation, ubiquitination, and other PTMs
-
Functional consequences of genomic mutations on protein abundance or activity
-
Alterations in protein complexes and interactomes
These insights enable mechanistic interpretation of tumor biology beyond what genome-level data can provide.
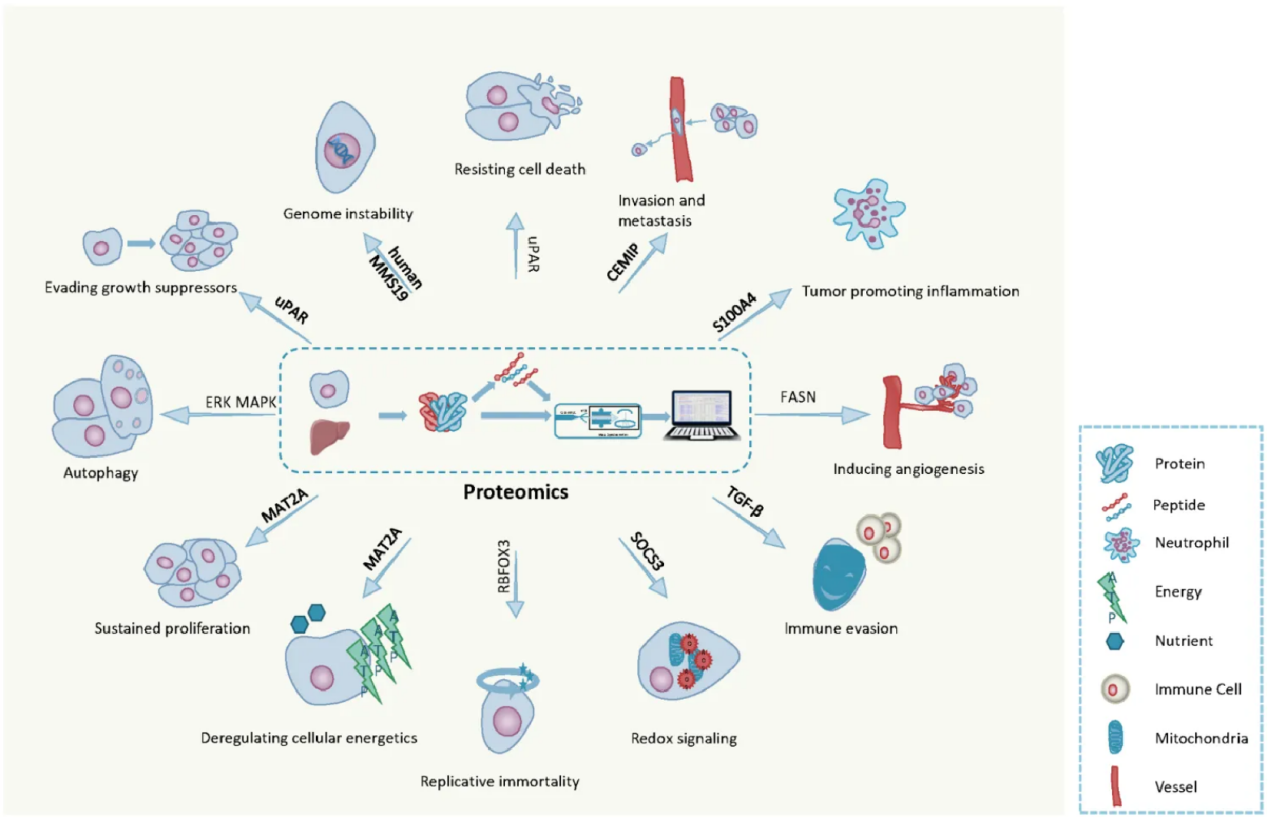
Su, M. et al. Cancers (Basel). 2021.
Figure 1. Cancer Proteomics Helps Reveal the Mechanisms of Cancer Development
2. Advancing Molecular Diagnosis
Proteins serve as direct readouts of disease state. Cancer proteomics supports:
-
Identification of diagnostic biomarkers from tissues, plasma, serum, urine, saliva, or exosomes
-
Stratification of disease subtypes based on proteomic signatures
-
Discovery of progression-associated markers for early detection
Quantitative proteomics and targeted MS refine diagnostic accuracy with high sensitivity and specificity.
3. Biomarker Discovery and Validation
Proteomics enables systematic discovery of:
-
Predictive biomarkers for treatment response
-
Prognostic markers associated with survival outcomes
-
Therapy-induced proteomic changes for monitoring clinical efficacy
Targeted MS workflows such as PRM, SRM, and MRM allow precise validation and quantification of candidate biomarkers across cohorts.
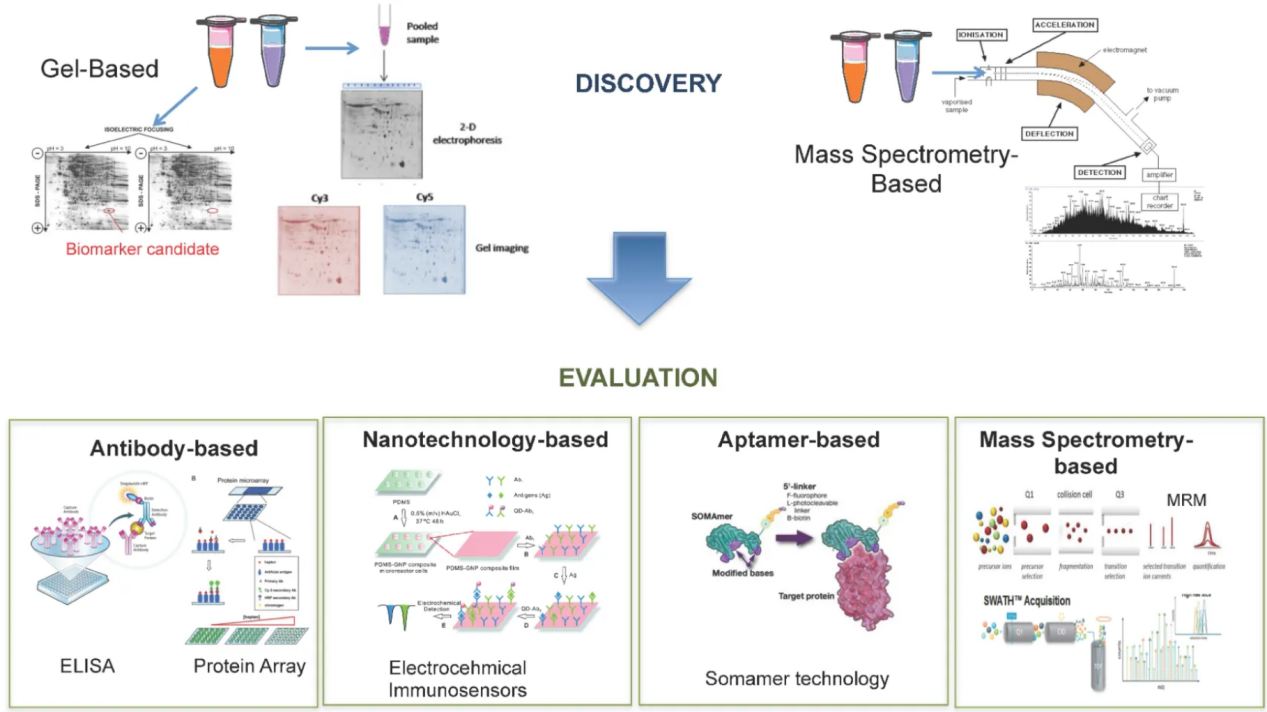
Tonry, C. L. et al. Diagnostics (Basel). 2016.
Figure 2. Proteomic Technology for Discovery and Evaluation of Protein Biomarkers
4. Drug Target Identification and Mechanistic Pharmacology
Proteomics is a powerful tool for therapeutic discovery through:
-
Mapping aberrant signaling networks to reveal targetable nodes
-
Identifying protein interactors, protein complexes, and ligand-binding proteins
-
Quantifying protein abundance changes upon drug treatment
-
Pharmacodynamic profiling and pathway-level response evaluation
Advanced chemoproteomics strategies can also identify covalent drug targets, off-target interactions, and ligandable hotspots.
Service at MtoZ Biolabs
MtoZ Biolabs integrates multiple analytical strategies to offer a full spectrum of proteomics capabilities for cancer research:
1. High-Resolution LC-MS/MS Discovery Proteomics
-
Deep proteome profiling with Orbitrap and timsTOF platforms
-
Data-dependent acquisition (DDA) and data-independent acquisition (DIA)
-
Comparison analysis across cancer phenotypes or treatment conditions
2. Quantitative Proteomics
-
TMT and iTRAQ multiplexed proteomics
-
SILAC for cell culture-based mechanistic studies
-
Label-free quantification (LFQ) with high reproducibility
3. PTM Profiling
Comprehensive identification and quantification of protein post-translational modifications, including but not limited to: Phosphorylation, Ubiquitination, Acetylation, Glycosylation, Methylation, Oxidation and redox modifications
These analyses reveal signaling activation, epigenetic regulation, and adaptive stress pathways in tumors.
4. Targeted Proteomics
PRM, SRM, and MRM for:
-
Clinical biomarker validation
-
Therapeutic monitoring
-
Quantitative assessment of low-abundance proteins
5. Subcellular Proteomics
Characterization of cancer-relevant compartments such as:
-
Membrane proteomes
-
Secretomes and exosomes
-
Mitochondrial proteomes
-
Nuclear and chromatin-associated proteomes
6. Bioinformatics Analysis
-
Data Quality Assessment
-
Multivariate PCA Analysis
-
Protein Statistical Analysis: Venn Diagram, Volcano Plot
-
Functional Annotation: GO Annotation, KEGG Annotation, COG Annotation
-
Clustering Analysis: Hierarchical Clustering, K-Means Clustering
-
Network Analysis: STRING Analysis
Analysis Workflow
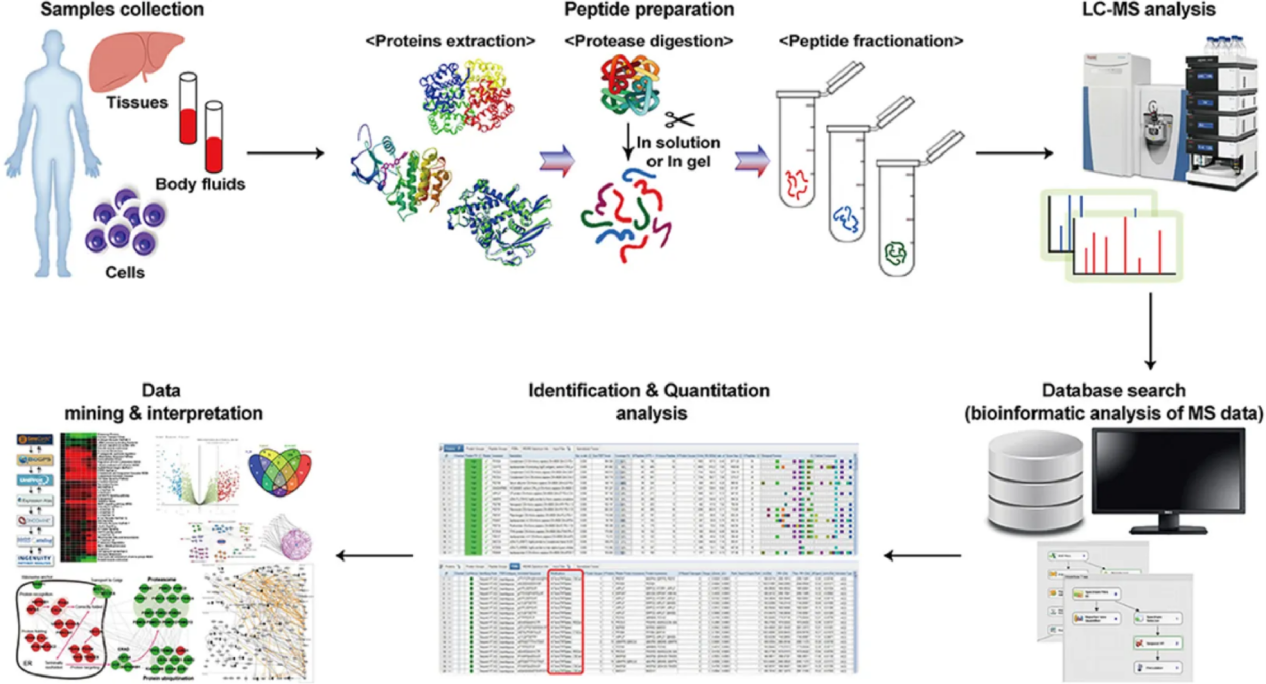
Kwon, Y. W. et al. Front Med (Lausanne). 2021.
Figure 3. Workflow of Cancer Proteomics Service
Why Choose MtoZ Biolabs
1. Advanced Platform
MtoZ Biolabs established an advanced cancer proteomics analysis platform, guaranteeing reliable, fast, and highly accurate analysis service.
2. High-Throughput Workflows
Automated sample handling, standardized LC-MS acquisition, and validated analytical pipelines offer consistent results across large cohorts.
3. Expertise in Oncology Proteomics
Our scientific team specializes in cancer proteomics, signaling pathway analysis, PTM biology, chemoproteomics, and quantitative methods such as DIA and PRM.
4. Customized Experimental Design
Every project is tailored to biological questions and sample constraints, ensuring maximum interpretability and research impact.
5. Secure, Traceable Data Delivery
All results are delivered with traceable QC metrics, raw files, processed data tables, and interpretive reports.
FAQ
Q1: What types of samples are suitable?
We support a wide range of cancer related samples, including
-
Fresh frozen or FFPE tumor tissues and biopsies
-
Paired tumor and adjacent normal tissues
-
Blood derived samples such as plasma, serum, and PBMCs
-
Cultured cancer cell lines
-
Patient derived organoids and xenograft models
If you have special or limited material, we can assess feasibility and optimize a suitable workflow.
Q2: How should I prepare my samples?
- Ensure samples are freshly prepared or snap-frozen.
- Avoid detergents, salts, and reducing agents that may interfere with LC-MS/MS analysis.
- Store samples at –80 °C before shipment and avoid repeated freeze thaw cycles.
- Transport samples on dry ice to maintain stability during delivery.
Detailed requirements and recommendations can be found in the Sample Submission Guidelines for Proteomics.
Q3: What is the service general workflow?
Q4: What data formats are provided?
- Raw data files in instrument specific formats
- Peptide and protein identification and quantitative result tables in .xlsx or .csv
- Statistical and bioinformatics outputs such as differential expression, pathway enrichment, and protein networks in spreadsheet formats
- Publication ready figures, for example heatmaps, volcano plots, PCA and clustering plots, in high resolution .png or .tiff
- A structured PDF report summarizing workflow, QC metrics, key findings, and biological interpretation
- Additional formats or customized deliverables can be provided on request to match your analysis or submission requirements.
Start Your Project with MtoZ Biolabs
Whether your objectives involve disease mechanism discovery, diagnostic signature development, therapeutic target identification, or translational biomarker validation, our team provides end-to-end support from experimental design through data interpretation.
To begin your project or discuss customized cancer proteomics solutions, contact the MtoZ Biolabs team today.
