MtoZ Biolabs provides Metaproteomics Services to characterize the functional protein landscape of complex microbial communities across clinical, environmental, and industrial settings. Our metaproteomics services use advanced 4D label-free quantitative proteomics to profile community structure, function, and dynamics at the protein level.
What Is Metaproteomics?
Metaproteomics is the large scale analysis of all proteins expressed by a mixed microbial community within a given environment, such as gut contents, soil, water, biofilms, or bioreactors. Instead of focusing on single isolates, metaproteomics captures the collective proteome of bacteria, archaea, fungi, viruses, and host associated proteins present in a sample.
By identifying and quantifying proteins in situ, metaproteomics reveals which metabolic pathways are active, how microbes interact with each other and with the host, and how community function responds to stimuli such as diet, drugs, pollutants, or environmental change. It links microbial composition to real biochemical activity, providing insight into energy metabolism, nutrient cycling, signaling, virulence, stress responses, and resistance mechanisms. This makes metaproteomics a powerful tool for microbiome research, environmental monitoring, agricultural improvement, industrial fermentation optimization, and translational studies in human and animal health.
Metaproteomics Service at MtoZ Biolabs
MtoZ Biolabs provides end to end Metaproteomics Service built on a 4D-LFQ platform that combines high resolution LC MS, ion mobility separation, and label free quantification. This 4D workflow improves separation of co eluting peptides in complex community samples, increases proteome depth, and enhances quantitative robustness for low abundance proteins.
You only need to define your study objectives and send us your samples. MtoZ Biolabs handles microbial protein extraction from challenging matrices, enzymatic digestion, peptide cleanup, MS acquisition, database search with microbiome focused resources, and downstream bioinformatics. We deliver taxonomic and functional annotations, differential protein and pathway profiles, and clear reports that link microbial community activity to your biological or environmental questions.
Workflow of Metaproteomics Service
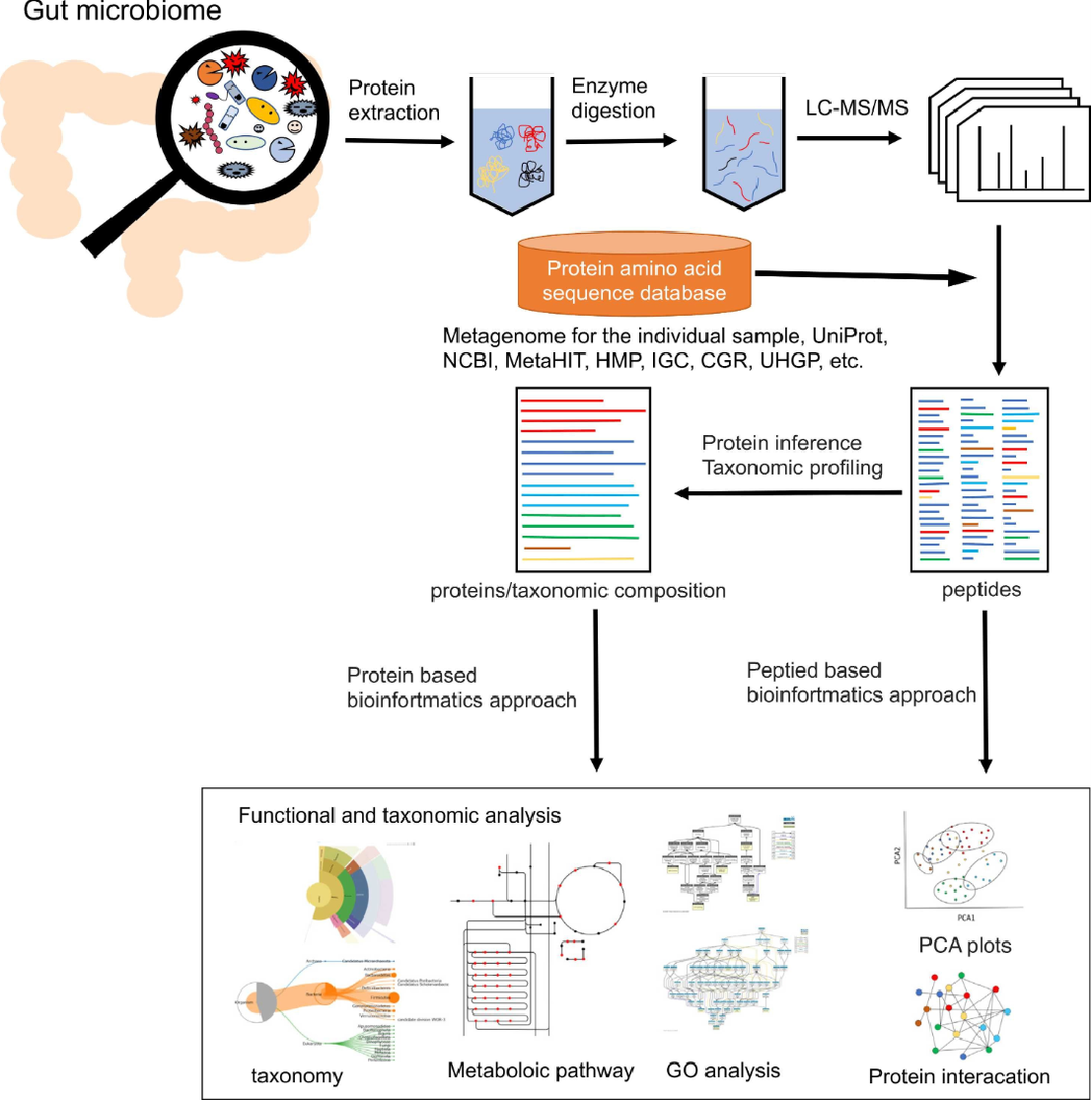
Miura, N. et al. Comput Struct Biotechnol J. 2023.
Figure 1. Metaproteomics Procedure
Why Choose MtoZ Biolabs
1. Customizable Service
We have dedicated protocols for complex community samples, from protein extraction to data interpretation, tailored to microbiome and environmental matrices.
2. Advanced Platform
Our 4D-LFQ Metaproteomics platform improves separation and sensitivity, enabling deeper coverage and more robust quantification of low abundance proteins in highly complex samples.
3. Experience with Diverse Sample Types
MtoZ Biolabs routinely handles feces, soil, sludge, water derived biomass, biofilms, plant associated samples, and animal associated microbiomes, with matrix specific optimization.
4. Flexible Study Design and Scaling
Our metaproteomics services support small exploratory studies, large multicenter cohorts, time course experiments, and industrial monitoring programs.
5. Transparent Communication and Support
From study design to result delivery, our scientific team works closely with you, providing clear updates, technical explanations, and follow up discussion as needed.
Applications of Metaproteomics Service
Metaproteomics Service at MtoZ Biolabs supports diverse research areas where understanding microbial community function is essential.
1. Human and Animal Microbiome Research
Study gut, oral, skin, respiratory, and reproductive tract microbiomes to link functional changes to diet, disease, drugs, and host physiology.
2. Environmental and Ecosystem Studies
Analyze soil, sediment, marine, freshwater, and wastewater communities to understand nutrient cycling, pollutant degradation, and ecosystem responses to environmental change.
3. Agriculture and Plant Associated Microbiomes
Investigate rhizosphere, phyllosphere, and endophytic microbiomes to explore plant growth promotion, stress tolerance, nutrient uptake, and disease suppression.
4. Industrial and Bioprocess Microbiology
Optimize fermentation, bioreactors, and bioenergy systems by monitoring community composition, metabolic activity, and process stability at the protein level.
5. Antimicrobial Resistance and Pathogenicity
Identify proteins linked to virulence, toxin production, stress adaptation, and resistance pathways in pathogenic or opportunistic microbial populations.
FAQ
Q1: What types of samples are suitable?
We accept a wide range of microbiome and environmental samples, including but not limited to:
- Fecal and intestinal content samples
- Soil, sediment, and sludge
- Water and wastewater microbiomes
- Biofilms and bioreactor communities
- Skin, oral, and other host associated microbiome samples
- Cultured microbial consortia and isolated community protein extracts
If you are unsure whether your sample type is suitable, you can contact us for a quick feasibility check.
Q2: How should I prepare my samples?
To ensure high quality metaproteomic data, we recommend:
- Use fresh or snap frozen samples; avoid repeated freeze-thaw cycles
- Collect samples in clean, low protein binding tubes or containers
- For most matrices, provide at least 0.5-1 g (solid) or 0.5-2 mL (liquid) when possible
- Store samples at -80°C and ship on dry ice
Detailed requirements and recommendations can be found in the Sample Submission Guidelines for Proteomics.
Q3: What is the service general workflow?
Q4: What data formats are provided?
- Raw data files in instrument specific formats
- Peptide and protein identification and quantification tables in Excel or CSV
- Taxonomic and functional annotation results, including pathway or GO summaries, in spreadsheet form
- Key visual outputs such as heatmaps, volcano plots, PCA, and bar charts in high resolution PNG or TIFF
- A structured PDF report describing workflow, QC metrics, main findings, and biological interpretation
Additional customized outputs or specific data formats can be provided upon request.
Start Your Project with MtoZ Biolabs
If you are planning a microbiome or community level study and want to connect microbial composition with real biochemical activity, Metaproteomics Service at MtoZ Biolabs can provide a complete, data rich solution from experimental design through to interpretable results.
Free project evaluation, welcome to learn more details!
