MtoZ Biolabs uses a high-resolution mass spectrometry platform and has launched the SRM/MRM targeted proteomics analysis service to perform highly sensitive and quantitative analysis of specific proteins. This service is widely applied in proteomics research, biomarker screening, and functional studies, providing precise quantitative data support for research.
Overview
SRM/MRM (selected/multiple reaction monitoring) is a type of targeted quantitative analysis technique based on triple quadrupole mass spectrometry. By selecting specific precursor ions and monitoring their characteristic fragment ions, it enables highly specific and highly sensitive detection of target peptides or proteins. Its principle relies on the sequential filtering and quantification capability of the triple quadrupole, allowing accurate tracking of abundance changes of specific molecules in complex backgrounds. SRM/MRM is widely used for targeted protein quantification and feature verification and is an important method for precise analysis of specific protein indicators.
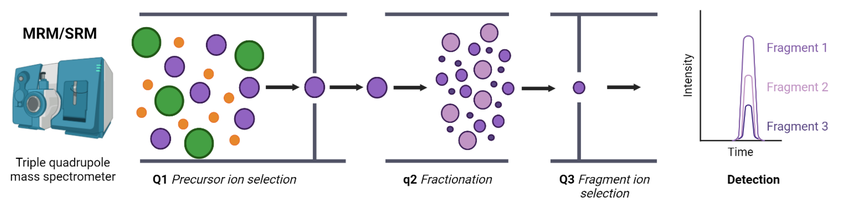
Guerrero, L N. et al. Metabolites, 2022.
Figure 1. Multiple/Selected Reaction Monitoring (MRM/SRM) Targeted Proteomics.
SRM/MRM Targeted Proteomics Analysis Service at MtoZ Biolabs
MtoZ Biolabs applies advanced SRM/MRM targeted mass spectrometry technology to provide high-sensitivity and highly reproducible quantitative proteomics analysis services for clients.
- Accurate quantification of target proteins
- Reliable detection of low-abundance proteins in complex systems
- Targeted quantification of common post-translationally modified proteins
- Quantitative comparison of protein abundance across multiple sample groups
Our analysis offers an in-depth understanding of the quantitative characteristics of target proteins, enabling researchers to obtain more reliable validation data and strengthen the conclusions of proteomics studies.
Workflow of SRM/MRM Targeted Proteomics Analysis Service
1. Sample Processing
Perform protein extraction, enzymatic digestion, purification, and stabilization to ensure that peptide quality meets the requirements of targeted analysis.
2. Target Selection and Method Development
Select signature peptides and ion pairs based on the target proteins and establish monitoring methods with high specificity and stability.
3. Mass Spectrometry Acquisition
Use a triple quadrupole platform to conduct SRM/MRM acquisition, accurately capturing signals of target peptides for highly selective detection.
4. Data Analysis
Perform statistical processing of ion-pair peak areas, retention times, and signal consistency to complete quantitative calculations and quality assessment.
5. Result Output
Provide standardized quantitative results, chromatographic and mass spectrometric images, and descriptions of experimental methods and parameters to generate a complete analysis report.
Why Choose MtoZ Biolabs?
- High Selectivity: Ensures target signal specificity through monitoring of characteristic precursor-product ion pairs
- High Sensitivity: Captures low-abundance target proteins or peptides within complex matrices
- High Throughput: Monitors multiple target peptides simultaneously to achieve parallel multi-indicator analysis
- Customized Analytical Solutions: Flexibly adjusts targets, parameters, and quantitative strategies based on research objectives
- One-Stop Support: Covers the complete targeted proteomics workflow from study design to final reporting
Applications of SRM/MRM Targeted Proteomics Analysis Service
1. Biomarker Discovery and Validation
By performing quantitative validation of candidate protein biomarkers, the stability and reproducibility under different samples or conditions can be evaluated.
2. Signal Pathway Studies
By monitoring abundance changes of key proteins or phosphorylation states in signaling pathways, the construction of regulatory networks can be supported.
3. Environmental Response Molecule Monitoring
Analyzes the expression levels of key proteins under changes in temperature, nutrients, or stress conditions to reveal adaptive response characteristics.
4. Metabolism-Related Protein Analysis
By evaluating the abundance of key regulatory proteins under different metabolic states, the understanding of metabolic regulatory processes can be assisted.
5. Quality Control and Batch Consistency Analysis
Used to assess the consistency of target proteins across production, expression, or experimental batches to ensure robust and reliable data.
Deliverables
- Comprehensive Experimental Details
- Materials, Instruments, and Methods
- Protein Quantification Results Table
- Mass Spectrometry Images and Visualization Charts
- Bioinformatics Analysis
- Raw Data Files
FAQ
Q1: What types of samples are suitable?
A1: Suitable samples include soluble, clarified, and relatively high-purity protein preparations, and multiple sources are accepted, including cell lysates, tissue extracts, biofluid samples, and immuno-enriched target proteins or peptides. Samples should minimize salts, surfactants, and complex matrix interference to ensure stability and accuracy in mass spectrometry-based quantification.
Q2: What is the service general workflow?
A2:
Q3: What data formats are provided?
A3: MtoZ Biolabs offers standardized data formats, including:
- Target protein quantification results table (XLSX/CSV)
- Mass spectrometry spectra and visualization charts (PNG/TIFF)
- Method parameter files (including chromatographic conditions and MRM/SRM transition lists)
- Comprehensive analysis report (PDF)
- Raw mass spectrometry data files (instrument-compatible formats)
If special analytical requirements exist, data formats can be customized according to project specifications.
Q4: How should I prepare my samples?
A4: To ensure optimal identification results, it is recommended to prepare samples in the following aspects:
- Purity: Samples should be free of precipitates and visible turbidity, with minimized salts, lipids, and surfactant residues
- Storage: For short-term storage, store at 4°C; for long-term storage, it is recommended to freeze at -80°C; avoid repeated freeze-thaw cycles
- Transportation: Use sealed, leak-proof containers and cold-chain shipping (ice packs/dry ice)
- Additional Information: Provide sample origin, buffer composition, and target protein/peptide information to support optimized method development and quantification strategies
For more information, please refer to Sample Submission Guidelines for Proteomics, Sample Submission Guidelines for Metabolomics.
Start Your Project with MtoZ Biolabs
Contact us to discuss your experimental design or request a quote. Whether you are focused on precise quantification of specific proteins or investigating key molecular changes within complex systems, we can provide high-sensitivity and high-confidence targeted analysis support.
