MtoZ Biolabs launches the Quantitative PTM Omics Service to leverage high-resolution mass spectrometry platforms and integrated multi-omics strategies to quantitatively profile post-translational modifications. This service helps research and industry users accurately capture regulatory signals, dynamic modification changes, and functionally important sites, providing high-quality data for mechanism studies, target discovery, and biomarker identification.
What is Quantitative PTM Omics
Quantitative PTM Omics refers to a comprehensive, systematic, and quantitative analytical approach for post-translational modifications within the proteomics framework. PTMs include phosphorylation, acetylation, methylation, ubiquitination, glycosylation, and others, all of which play important roles in signal transduction, metabolic regulation, protein stability, and cell fate determination.
Through high-resolution LC-MS/MS, PTM-specific enrichment, DIA strategies, or labeling-based quantitative methods, Quantitative PTM Omics enables site-level identification, abundance measurement, and cross-condition comparison. Compared with traditional protein expression analysis, it reveals dynamic regulatory changes and functional nodes within cellular networks.
Quantitative PTM Omics Service at MtoZ Biolabs
MtoZ Biolabs provides Quantitative PTM Omics Service focusing on high sensitivity quantification of diverse PTM types, including:
• Parallel profiling of multiple PTMs such as phosphorylation, acetylation, methylation, ubiquitination, glycosylation, and nitration
• Site-level identification using PTM enrichment coupled with high-resolution MS
• DIA or labeling strategies such as TMT for quantitative comparison
• Functional annotation, pathway mapping, and regulatory network analysis
• Optional integration with multi-omics data, including transcriptomics, proteomics, and metabolomics
Table 1. Detectable PTM Proteomics at MtoZ Biolabs
| Ubiquitinated Proteomics Analysis Service | Nitrosation Omics Service | Lipid-Modified Proteomics Service | 4D Post-translational Modifications (PTMs) Proteomics Service | 4D-Methylation Proteomics Service |
| Acetylomics Analysis Service | S-Nitrosoproteomics Service | Propionylomics Service | 4D-Ubiquitination Proteomics Service | 4D-Glycosylation Proteomics Service |
| High-Resolution Phosphoproteomics Service | Disulfidomics Service | Citrullinomics Service | 4D-Acetylation Proteomics Service | |
| Integrated Methylation Proteomics Service | Advanced Glycoproteomics Service | Meta-PTMomics Analysis Service | 4D-Phosphoproteomics Service |
Our service supports a wide range of sample types, including cell lines, tissues, biofluids, and model organisms.
Workflow of Quantitative PTM Omics Service
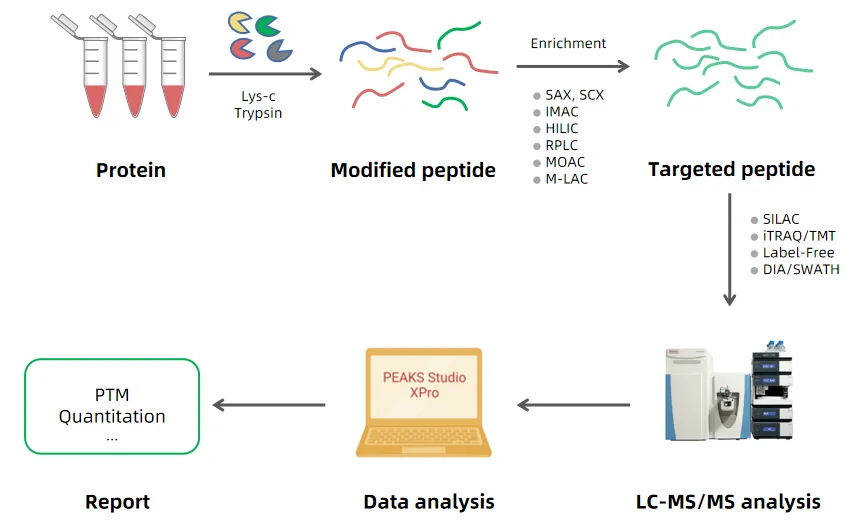
Why Choose MtoZ Biolabs
1. High-Resolution Mass Spectrometry Platforms
Orbitrap Fusion Lumos and Q Exactive HF systems provide high sensitivity, broad dynamic range, and excellent reproducibility.
2. Parallel Detection of Multiple PTM Types
Supports enrichment and quantification of diverse modifications suitable for complex biological regulation studies.
3. Multiple Quantitative Strategies
Supports multiple quantitative strategies, including label-free, TMT multiplexing, DIA, and PRM, allowing flexible selection of the optimal approach based on PTM type and experimental design.
4. Experienced Project and R&D Teams
Familiar with sample preparation challenges and common sources of interference to ensure reliable data quality.
Applications of Quantitative PTM Omics Service
1. Signal Transduction and Dynamic Regulatory Mechanisms
Analysis of kinase regulation, phosphorylation switches, and modification cascades.
2. Cell State and Environmental Response
Profiling PTM changes under oxidative stress, drug treatment, or metabolic perturbation.
3. Mechanistic Studies in Cancer, Metabolic Disease, and Immune Disorders
Identification of functional modification sites to support target discovery.
4. Drug Mechanism and Resistance Studies
Capturing PTM alterations associated with drug response to support pharmacodynamic evaluation.
5. Protein Degradation, Stability, and Interaction Network Analysis
Using ubiquitination, acetylation, and other PTMs to map key regulatory determinants of protein fate.
Deliverables
• PTM identification list with site and modification information
• Protein and peptide quantification tables
• Functional enrichment and pathway analysis results
• Summary of differential modifications and key regulatory nodes
• Raw MS data and method documentation
FAQs
Q1: What types of samples are suitable?
Suitable samples include cells, tissues, serum, plasma, model organism samples, and recombinant proteins, all of which can be used for multi type PTM quantification studies.
Q2: What is the service's general workflow?
Q3: What data formats are provided?
MtoZ Biolabs provides results in multiple standard formats to ensure compatibility with common analytical and visualization tools. Deliverables typically include:
• Raw data files generated by high-resolution LC-MS/MS platforms
• Processed data tables in CSV or Excel format containing identified proteins, peptides, and PTM sites
• Summary reports in PDF format, including analytical methods, quality control metrics, and major results
• High-resolution figures such as spectra, heatmaps, and pathway maps in TIFF or PNG
Additional formats or structured data outputs can be provided upon request to support specific research or publication needs.
Q4: How should I prepare my samples?
To ensure accurate and reproducible Quantitative PTM Omics results, MtoZ Biolabs recommends the following preparation guidelines:
1. Sample Type: Accepted samples include cultured cells, tissues, serum or plasma, microbial samples, and purified protein preparations.
2. Sample Purity: Samples should be free of excessive salts, detergents, lipids, or inhibitors that may interfere with enzymatic digestion or mass spectrometry ionization.
3. Protein Amount Requirement: Provide at least 200 µg of total protein. For PTM enrichment workflows, 500 µg or more may be required, depending on the modification type.
4. Storage and Shipping: Store samples at -80℃. Ship samples on dry ice using sealed, leak-proof containers to maintain sample stability.
5. Documentation: Include information such as sample origin, preparation method, treatment conditions, buffer composition, and the type of PTM analysis requested.
For more information, please refer to Sample Submission Guidelines for Proteomics and Sample Submission Guidelines for Metabolomics.
