MtoZ Biolabs provides DIA Quantitative Proteomics Analysis Service using data independent acquisition LC-MS/MS to deliver deep, reproducible, and high throughput protein quantification. This service supports discovery proteomics, biomarker screening, and large scale comparative studies across cells, tissues, and biofluids.
What Is DIA Quantitative Proteomics?
DIA (Data-Independent Acquisition) quantitative proteomics is a mass spectrometry strategy in which all ions within predefined m/z windows are systematically fragmented, and the resulting fragment ion signals are used for peptide identification and quantification. Compared with traditional data dependent acquisition (DDA), DIA generally provides more consistent peptide detection across runs, fewer missing values, and more reliable quantification for low to medium abundance proteins in complex samples.
Principle of DIA Quantitative Proteomics Analysis
In a classic DDA workflow, the instrument acquires an MS1 survey scan, then automatically selects the top N most intense precursor ions for MS/MS fragmentation. This intensity driven selection means that lower abundance or transient peptides may not be chosen in every run, so different samples can yield different sets of identified peptides. As a result, DDA data often show stochastic coverage and gaps in the quantitative matrix when many samples or conditions are compared.
By contrast, DIA divides the full m/z range into consecutive or overlapping isolation windows and cycles through them, fragmenting all precursors within each window in a systematic and repeated manner. The instrument generates a comprehensive fragment ion map across the LC gradient, capturing signals from both high and low abundance peptides. Peptides are then identified and quantified by targeted extraction of characteristic fragment ion traces, guided by spectral libraries or library free algorithms. This acquisition principle greatly reduces the stochastic behavior seen in DDA and enables DIA to deliver more complete, consistent, and robust quantitative proteomic data for downstream biological interpretation.
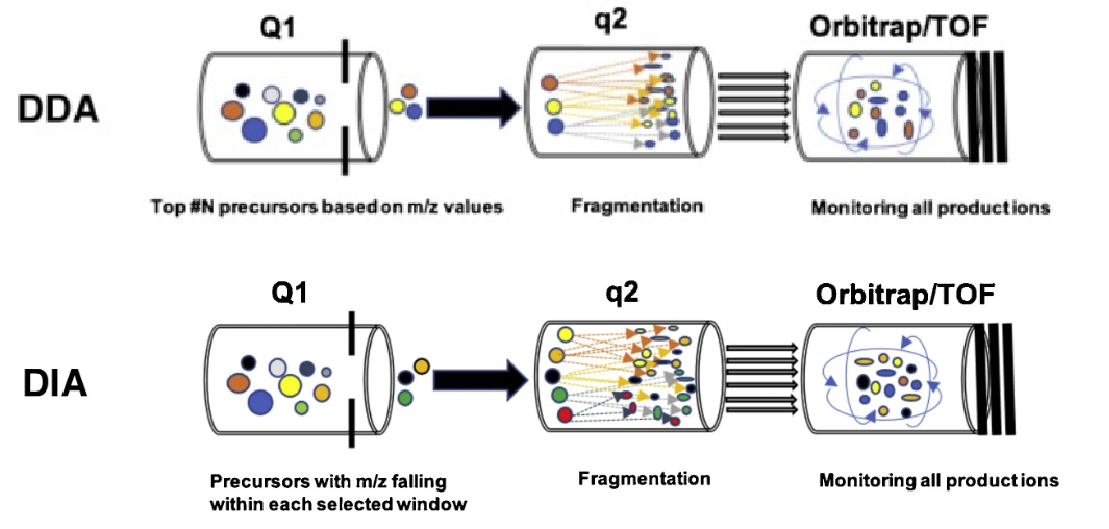
Li J. et al. Drug Discov Today Technol. 2021.
Figure 1. Principle of DDA and DIA
DIA Quantitative Proteomics Analysis Service at MtoZ Biolabs
MtoZ Biolabs offers DIA Quantitative Proteomics Analysis Service on high-resolution LC-MS platforms with optimized DIA acquisition schemes and analysis pipelines. The service is designed to support both moderate sized experiments and large clinical style cohorts.
We provide the following specialized DIA services:
- Direct DIA Service
Streamlined data-independent acquisition with machine learning to generate high-quality peptide libraries without the need for a pre-built spectral library.
- GPF-DIA Quantitative Proteomics Service
Leverage gas-phase fractionation to enhance sensitivity and reduce co-fragmentation, improving the detection of low-abundance proteins and increasing the depth of analysis.
- PCT-DIA Service
Combining pressure cycling technology with DIA to optimize protein extraction and fragmentation, enhancing reproducibility and consistency across samples.
- SWATH-MS Quantitative Proteomics Service
Using SWATH (Sequential Window Acquisition of All Theoretical Fragment Ions) for comprehensive, high-precision quantitative analysis of complex samples.
- 4D-DIA Quantitative Proteomics Analysis Service
Integrating ion mobility with DIA for deeper proteome coverage, improved quantification accuracy, and better resolution of co-eluting peptides.
- MSX-DIA Quantitative Proteomics Service
A multiplexed DIA approach that improves quantification accuracy and throughput by optimizing precursor ion selection, enhancing sensitivity and reducing interference.
You only need to define your study objectives and send us your samples. MtoZ Biolabs takes care of protein extraction, digestion, DIA LC-MS/MS acquisition, data processing, quantitative analysis, and interpretation, delivering a complete and ready to use data package.
Workflow of DIA Quantitative Proteomics Analysis Service
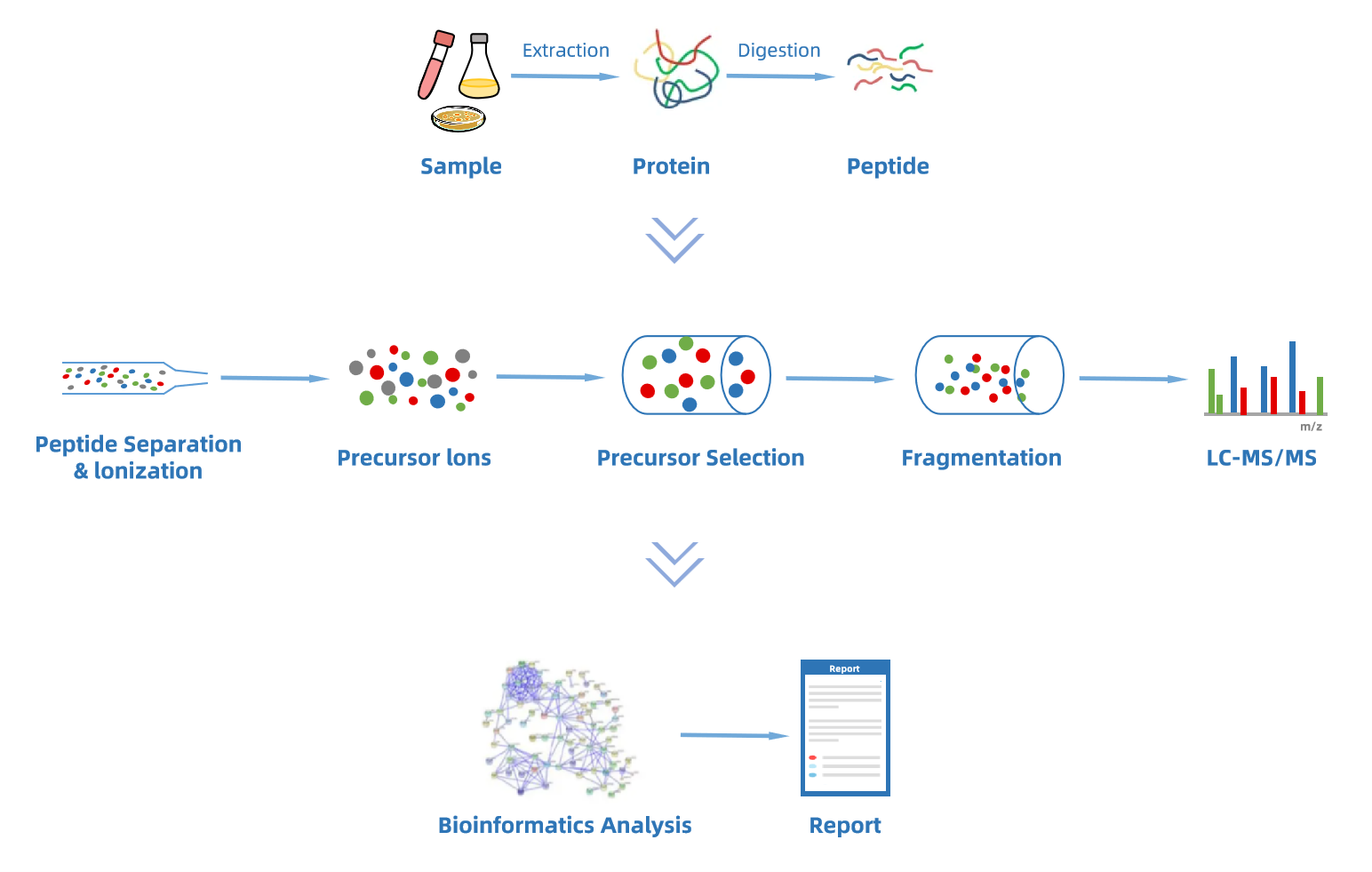
Why Choose MtoZ Biolabs
1. High-Resolution Platforms
We utilize advanced LC-MS/MS systems, including Thermo Fisher Orbitrap, to ensure high sensitivity and accuracy for complex proteomic analysis.
2. Comprehensive Proteome Coverage
Our DIA approach captures a broad range of peptides, including low-abundance proteins, providing a deep and unbiased snapshot of the proteome.
3. Flexible and Scalable
Our service supports a variety of experimental designs, from small-scale studies to large cohorts, offering tailored solutions for discovery, biomarker, and pathway analysis.
4. End-to-End Support
From experimental design to bioinformatics analysis, our team provides expert guidance and a seamless workflow, delivering high-quality, interpretable results.
Applications of DIA Quantitative Proteomics Analysis Service
1. Biopharmaceutical Research
DIA-based proteomics supports the discovery and validation of protein biomarkers, helping identify therapeutic targets, assess drug efficacy, and monitor therapeutic protein production processes in biopharmaceutical development.
2. Metabolic and Nutritional Studies
Quantifying protein changes in response to diet, metabolic imbalances, or environmental stressors, DIA proteomics helps uncover the molecular mechanisms behind metabolic diseases and informs nutrition-based interventions.
3. Toxicology and Environmental Health
By monitoring proteomic changes in response to environmental toxins or drug exposure, DIA proteomics helps assess toxicological impacts, identify biomarkers of exposure, and support safety evaluations for pharmaceutical and chemical industries.
4. Basic Biological Research
DIA proteomics is used to investigate cellular mechanisms such as signal transduction, gene expression regulation, and post-translational modifications, aiding in the understanding of basic cellular processes and their dysfunctions in disease.
FAQ
Q1: What types of samples are suitable?
We accept a variety of sample types for DIA Quantitative Proteomics, including:
- Cell pellets from adherent or suspension cell lines
- Fresh frozen tissues or biopsies
- Biofluids such as serum, plasma, cerebrospinal fluid (CSF), urine
- Primary cells from blood, bone marrow, or other tissues
- Preclinical or animal model samples for comparative studies
For unusual sample types or low-yield materials, please contact MtoZ Biolabs for customized preparation guidance.
Q2: How should I prepare my samples?
- Use clean, low binding tubes and avoid repeated freeze thaw cycles
- Snap freeze samples after collection and store at -80 °C
- Avoid strong detergents, very high salt, and large amounts of reducing agents
- Ship on dry ice with a clearly labeled sample list
Detailed requirements and recommendations can be found in the Sample Submission Guidelines for Proteomics.
Q3: What is the service general workflow?
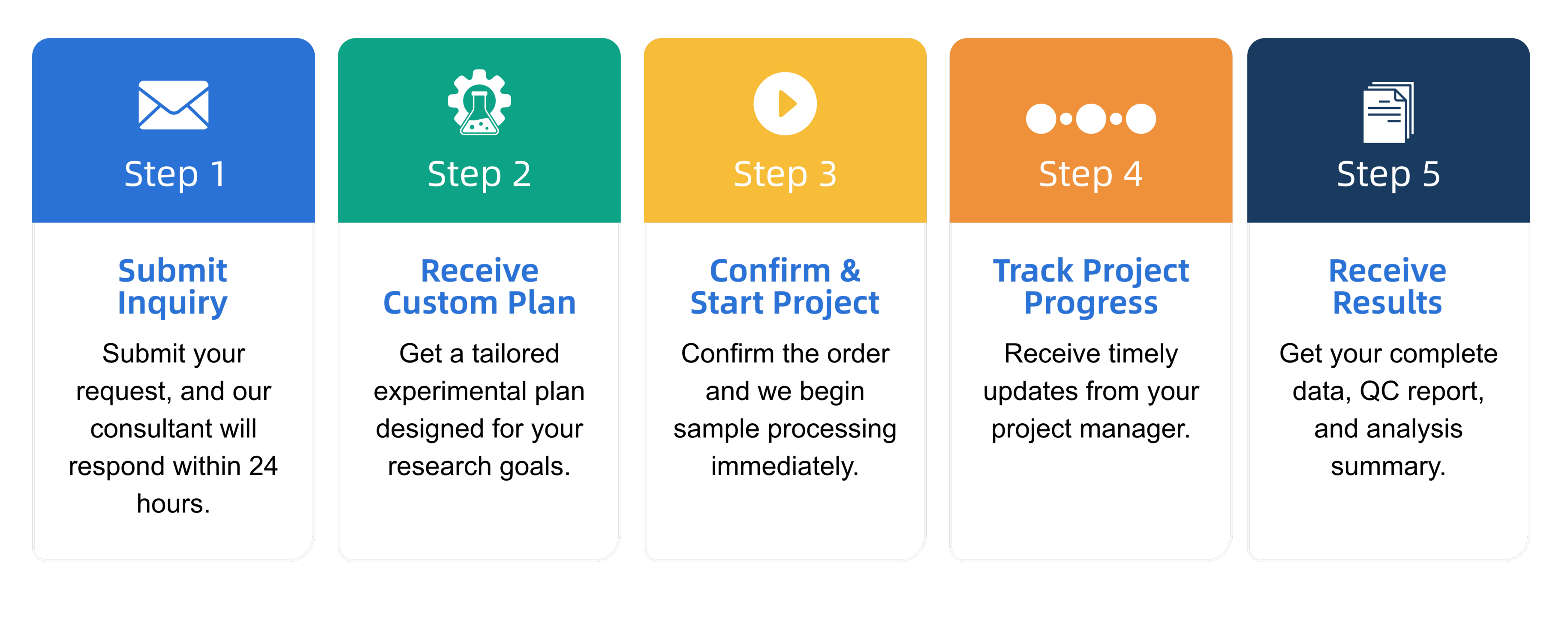
Q4: What data formats are provided?
- Raw LC-MS/MS files in instrument-specific formats.
- Quantitative peptide and protein tables in .xlsx or .csv format.
- Statistical analysis results, including fold change, p-values, and enrichment analysis, in spreadsheet format.
- High-resolution visual outputs such as heatmaps, volcano plots, and PCA plots in PNG or TIFF format.
- A comprehensive PDF report summarizing experimental design, analysis methods, QC metrics, and biological interpretations.
- Additional formats or custom analysis outputs can be provided on request.
Start Your Project with MtoZ Biolabs
Whether you are investigating disease mechanisms, identifying biomarkers, or conducting large-scale comparative studies, our DIA Quantitative Proteomics Analysis Service provides the data quality and biological insights you need to advance your research.
To learn more or discuss your project requirements, contact MtoZ Biolabs today. Our team is ready to assist you in designing and executing a proteomics workflow tailored to your specific needs.
