MtoZ Biolabs provides comprehensive Post-Translational Modification Proteome Profiling Service to support mechanism driven basic research, translational studies, and drug discovery projects. Using advanced LC-MS-based proteomics platforms, we help you characterize, quantify, and interpret diverse post-translational modifications across proteins and pathways in a wide range of biological systems.
What Is Post-Translational Modification Proteome?
Post translational modifications are chemical changes that occur on proteins after translation, such as phosphorylation, ubiquitination, acetylation, methylation, glycosylation, SUMOylation, and lipidation. These dynamic modifications regulate protein activity, localization, stability, complex formation, and signaling output, and they lie at the core of processes such as cell cycle control, immune responses, metabolism, and stress adaptation.
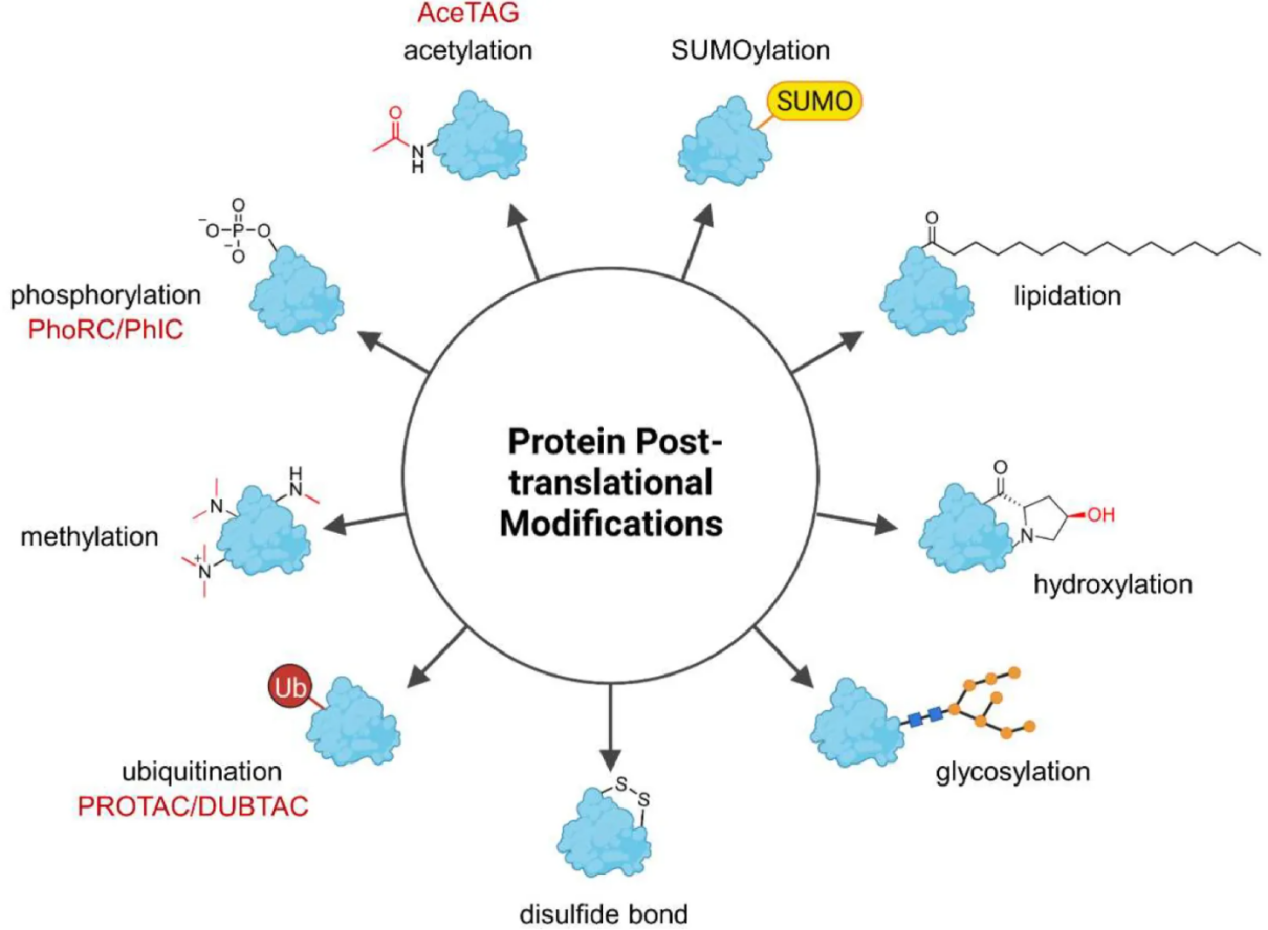
Heitel, P. Molecules. 2023.
Post-Translational Modification (PTM) proteome profiling is the systematic, large scale analysis of modified peptides and proteins in biological samples using mass spectrometry. Instead of measuring only total protein abundance, PTM profiling resolves which sites are modified, to what extent, and how those modifications change across conditions.
Post-Translational Modification Proteome Profiling Service at MtoZ Biolabs
By combining selective enrichment strategies with high–resolution LC–MS/MS and advanced quantitative methods, protein PTM profiling at MtoZ Biolabs delivers both site specific detail and global systems level insight.
1. PTM Qualitative Analysis Service
- Target Protein PTM Analysis
For hypothesis driven projects focused on a single protein or defined complex, we combine immunoprecipitation or affinity enrichment with high-resolution LC-MS/MS to map modification sites at amino acid resolution. This service characterizes which residues are modified, the coexistence of multiple PTMs on the same protein, and PTM patterns under different treatments, supporting mechanistic studies, mutant design, and target validation.
- PTM Omics Identification (Discovery Level)
For global PTM screening, we apply PTM specific enrichment strategies such as phosphopeptide, ubiquitinated peptide, acetylated peptide, or glycopeptide enrichment, followed by DDA or DIA-based LC-MS/MS. This module delivers large scale catalogs of modified peptides, confidently localized sites, and associated proteins, revealing pathway level regulation and providing candidate PTM events for downstream validation or targeting.
2. Quantitative PTM Omics Service
The Quantitative PTM Omics Service is designed to quantify PTM changes across conditions, time points, or dose groups with high reproducibility. Depending on project needs, we use label free DIA and 4D workflows, isobaric labeling strategies such as TMT or iTRAQ, or targeted PRM SRM panels built from discovery data. You receive normalized quantitative profiles of PTM sites and modified proteins, statistical comparisons between groups, and pathway and network level interpretations that distinguish PTM specific regulation from total protein abundance changes.
Workflow of Post-Translational Modification Proteome Profiling Service
1. Sample Receipt and Preparation
Biological samples are received under controlled conditions, logged, and processed with optimized lysis buffers and inhibitors to preserve labile PTMs.
2. Protein Extraction, Digestion, and PTM Enrichment
Total proteins are extracted, reduced, alkylated, and enzymatically digested into peptides. PTM specific enrichment strategies, such as IMAC, MOAC, lectin-based and HILIC methods, are then applied as required by the project.
3. LC-MS/MS Analysis
Enriched PTM peptides are separated by high performance nano LC and analyzed on high resolution mass spectrometry platforms using DDA, DIA, or targeted PRM, SRM methods to obtain high quality MS and MS/MS spectra.
4. Data Processing and Bioinformatics Analysis
Raw data are processed for peptide and site identification, PTM site localization, and quantitative comparison across groups. Downstream bioinformatics includes pathway and network analysis to interpret PTM regulation in a biological context.
5. Reporting and Scientific Support
MtoZ Biolabs delivers a comprehensive report including experimental workflow, QC metrics, PTM site tables, statistical results, and key biological insights, along with optional follow up consultation to support publication or subsequent validation experiments.

Why Choose MtoZ Biolabs
1. High Performance Mass Spectrometry Platforms
High resolution LC–MS configurations with DIA, DDA, 4D, and targeted acquisition modes to match your depth, throughput, and sensitivity requirements.
2. Site Level Precision and Rigorous QC
Emphasis on robust site localization, quantitative reproducibility, and comprehensive quality metrics throughout the workflow.
3. Flexible Study Designs
Support for small exploratory projects, multi-group comparison studies, and large cohort or time course designs.
4. Expert Data Analysis and Interpretation
Dedicated bioinformatics support for PTM-specific pipelines, including site centric analysis, pathway mapping, and integration with existing datasets.
5. Collaborative Project Management
Close communication from experimental design through reporting, with the goal of delivering datasets that are ready for direct biological interpretation and publication.
Applications of Post-Translational Modification Proteome Profiling Service
- Global profiling of phosphorylation, ubiquitination, acetylation, glycosylation and other PTMs to dissect signaling pathways and regulatory networks
- Characterizing disease related PTM changes in cancer, inflammatory disorders, neurodegeneration, metabolic and cardiovascular diseases
- Evaluating drug mechanism of action, target engagement and pathway modulation in preclinical and translational studies
- Discovering and prioritizing PTM-based biomarkers for patient stratification, prognosis and treatment response monitoring
- Comparing PTM landscapes across cell types, developmental stages, stress conditions or genetic perturbations
- Investigating PTM crosstalk, protein complex dynamics and pathway rewiring under physiological or pathological stimuli
FAQ
Q1: What types of samples are suitable?
We accept a wide range of PTM-compatible samples, including but not limited to:
- Cultured cells and primary cells
- Fresh or frozen tissues from human, animal, plant, or microbial sources
- Biofluids such as plasma, serum, CSF, urine, and cell culture supernatant
- FFPE tissues and tissue sections
If you have a special matrix or very limited amounts, you can contact us in advance for a feasibility evaluation.
Q2: How should I prepare my samples?
- Avoid strong detergents and chaotropic agents that are not MS compatible whenever possible
- Use protease and, when relevant, phosphatase or deubiquitinase inhibitors during lysis to preserve PTMs
- Snap freeze samples in liquid nitrogen and store at -80°C, minimizing freeze-thaw cycles
- Provide detailed information about sample type, number of conditions and replicates, and any previous treatments
Detailed requirements and recommendations can be found in the Sample Submission Guidelines for Proteomics.
Q3: What is the service general workflow?
Q4: What data formats are provided?
- Raw data files in instrument specific formats
- Processed peptide and protein identification and quantification tables in Excel or CSV
- PTM site level tables including positions, localization probabilities, and quantitative values when applicable
- Bioinformatics outputs such as differential analysis, motif analysis, and pathway or enrichment results in spreadsheet formats
- Key visualizations, for example heatmaps, volcano plots, PTM site distribution, and pathway diagrams in high-resolution image formats
- A structured PDF report summarizing workflow, quality control, main findings, and biological interpretation
Additional customized deliverables can be provided on request to match your downstream analysis or publication needs.
Start Your Project with MtoZ Biolabs
With MtoZ Biolabs as your partner, you can systematically explore the PTM landscape of your system of interest, connect modification patterns to function and phenotype, and drive your research or development programs forward with high-confidence proteomics data.
Free project evaluation, welcome to learn more details!
