MtoZ Biolabs employs advanced instrumentation and analytical platforms to provide the quantitative plant proteome analysis service for systematic quantitative analysis of protein expression in plant samples. This service is applicable to studies of plant growth and development as well as stress responses, providing reliable data support for related proteomics analyses.
Overview
Plant proteomics is a research field that focuses on plant samples and aims to systematically identify and analyze protein composition, expression levels, and their dynamic changes. As an important link between the genome and phenotype, the proteome more directly reflects plant physiological states and regulatory processes. Quantitative plant proteomics helps reveal dynamic changes in protein expression under different conditions and has broad applications in plant growth and development, stress response studies, metabolic regulation, and functional gene research.
Quantitative Plant Proteome Analysis Service at MtoZ Biolabs
MtoZ Biolabs provides a comprehensive range of quantitative plant proteomics analysis services to meet the needs of different research stages and experimental designs. The service offerings include, but are not limited to, the following analysis types:
1. Label-Free Quantitative Plant Proteomics Analysis
Relative quantification is performed based on mass spectrometry signal intensity without the use of chemical or isotopic labeling, making this approach suitable for large-scale plant samples and multi-condition comparative studies.
2. SWATH Quantitative Plant Proteomics Analysis
A wide mass window-based systematic acquisition strategy is applied to achieve stable and reproducible quantitative results, and is well suited for cross-batch or long-term project analyses.
3. DIA Quantitative Plant Proteomics Analysis
Data-independent acquisition is used to comprehensively record peptide information, reducing acquisition randomness and improving quantitative consistency and proteome coverage.
4. Label-Based Quantitative Plant Proteomics
Chemical or isotopic labels are used to distinguish different samples, enabling multi-sample quantitative comparison within a single mass spectrometry run.
5. TMT Quantitative Plant Proteomics Analysis
Supports multiplexed sample analysis and is suitable for high-precision quantitative studies with strong comparability under complex experimental designs.
6. iTRAQ Quantitative Plant Proteomics Analysis
Uses isobaric tags to achieve quantitative analysis of multiple samples, and is commonly applied to compare protein expression differences across treatments, cultivars, or phenotypes.
7. 2D-DIGE Quantitative Plant Proteomics Analysis
Combines two-dimensional gel electrophoresis with differential fluorescent labeling to visually display changes in protein abundance and overall proteome profile differences.
Workflow of Quantitative Plant Proteome Analysis Service
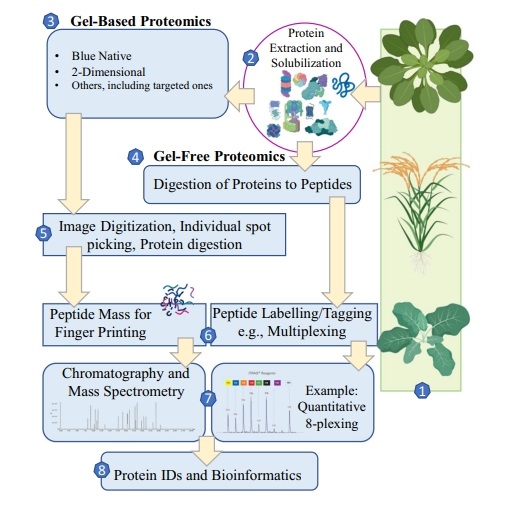
Ahmad, J. et al. Environmental Chemistry Letters, 2022.
Figure 1. Quantitative Analysis Workflow of Plant Proteome.
Why Choose MtoZ Biolabs?
- Diverse Quantitative Strategies: Cover multiple plant protein quantification approaches, including label-free, DIA, TMT, and iTRAQ
- Stable and Reliable Data: Standardized workflows ensure comparability of results across different batches and samples
- Strong Adaptability to Complex Samples: Suitable for plant samples with diverse tissue types and complex backgrounds
- High-Throughput Analysis Capability: Supports parallel analysis of multiple samples to improve overall research efficiency
- Customized Analysis Design: Enables flexible selection of appropriate quantitative strategies based on research objectives
Applications of Quantitative Plant Proteome Analysis Service
1. Plant Growth, Development, and Organ Differentiation Studies
Used to systematically analyze changes in protein expression across different growth stages and plant tissues or organs, supporting the understanding of growth and developmental regulation.
2. Stress Response and Environmental Adaptation Studies
Support the characterization of protein response patterns and adaptive processes in plants under drought, salinity stress, temperature variation, or pathogen infection.
3. Metabolic Regulation and Physiological Process Studies
Applied to investigate expression changes and regulatory relationships of proteins involved in photosynthesis, energy metabolism, and secondary metabolism.
4. Signal Transduction and Regulatory Network Studies
Assist in analyzing expression changes of key proteins within plant hormone signaling and stress-related pathways, supporting the construction and interpretation of regulatory networks.
5. Plant-Environment Interaction Studies
Support the analysis of molecular interactions between plants and environmental factors, helping to elucidate plant response strategies to external condition changes.
Deliverables
- Comprehensive Experimental Details
- Materials, Instruments, and Methods
- Tables of Quantitative Analysis Results for Plant Proteins
- Relevant Mass Spectrometry Spectra and Visualized Figures
- Bioinformatics Analysis
- Raw Data Files
FAQ
Q1: What types of samples are suitable?
A1: The quantitative plant proteome analysis service is suitable for well-characterized plant samples in good condition, including different tissues and organs such as roots, leaves, stems, flowers, and seeds, as well as plant materials from different treatments or developmental stages. Samples should maintain good integrity and avoid obvious degradation or severe interference from contaminants to meet the requirements of quantitative proteomics analysis.
Q2: What is the service general workflow?
A2:
Q3: What data formats are provided?
A3: MtoZ Biolabs provides multiple types of standardized deliverables, including:
- Protein identification and quantitative result tables (Excel/CSV)
- Protein differential analysis and related statistical files
- Visualized result figures (PNG/TIFF)
- Raw and processed mass spectrometry data files
- Comprehensive analysis report (PDF)
If special analytical requirements exist, data formats can be customized according to project specifications.
Q4: How should I prepare my samples?
A4: Sample preparation is recommended in the following aspects:
- Sample Purity: Remove soil, polysaccharides, pigments, and other impurities as much as possible, and avoid obvious protein degradation.
- Sample Storage: Samples should be stored at low temperature in a timely manner; long-term storage is recommended at -80℃ and repeated freeze-thaw cycles should be avoided.
- Sample Transport: Samples should be transported under low-temperature and sealed conditions to ensure stability.
- Additional Information: Providing sample source, tissue type, treatment conditions, and experimental grouping information will help optimize the analysis strategy.
For more information, please refer to Sample Submission Guidelines for Proteomics, Sample Submission Guidelines for Metabolomics.
Start Your Project with MtoZ Biolabs
Contact us to discuss your experimental design or request a quote. Whether you are exploring changes in plant protein expression or conducting related functional studies, we can provide you with professional and reliable analytical support.
