MtoZ Biolabs utilizes advanced instrumentation and analytical platforms to launch the large-scale protein identification service which provides systematic identification and analysis of large-scale protein compositions in complex samples. This service is widely used in proteomics research, drug development, and biomarker discovery, providing reliable data support for a deeper understanding of protein expression features in biological systems.
Overview
Proteins are the fundamental molecules of life, formed by the specific sequence of amino acids, and perform key functions such as catalysis, growth, signal transduction, and structural support. Systematic protein identification clarifies the protein composition and characteristic variations in samples, making it a crucial aspect of proteomics research. Large-scale protein identification has been widely applied in proteomics, disease research, biomarker discovery, and drug development, providing reliable data support for related studies.
Large-Scale Protein Identification Service at MtoZ Biolabs
1. SDS-PAGE Spot and Band Protein Identification
By extracting the target protein from SDS-PAGE spots or bands, enzymatically digesting, and performing mass spectrometry analysis, the identity of a single or small amount of protein can be quickly confirmed. This is suitable for accurate identification of band-level or spot-based proteins.
2. Protein Mass Spectrometry Identification
Using the high-resolution LC-MS/MS platform, peptide fragment information is obtained, and protein identity is confirmed through database comparison. This method is suitable for routine protein identification needs in various sample types.
3. Shotgun Protein Identification
After directly digesting complex protein mixtures into peptides, large-scale mass spectrometry analysis is performed. This approach can identify thousands of proteins in a single experiment, providing high-throughput support for systematic protein composition analysis.
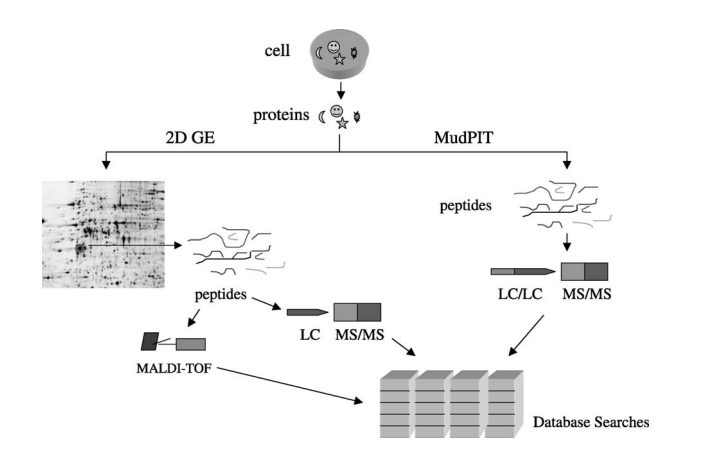
Lin, D Y. et al. BBA - Protein and Proteomics, 2003.
Figure 1. Schemes for Large-Scale Protein Identification Methods.
Workflow of Large-Scale Protein Identification Service
1. Sample Preparation
The protein sample is dissolved, desalted, and pre-purified to ensure it is suitable for mass spectrometry analysis.
2. Protein Enzyme Digestion
Appropriate proteases are used to digest the protein into peptides, providing the foundation for subsequent analysis.
3. Mass Spectrometry Analysis
High-resolution mass spectrometry technology is employed to acquire peptide fragment ion signals, which are then used for protein identification.
4. Data Processing and Analysis
Peptide fragment data is compared against a database to deduce the protein sequence and identify the types of proteins present.
5. Result Output
A detailed identification report is provided, including protein information, amino acid sequences, and related analysis data.

Figure 2. The Workflow of Protein Identification.
Why Choose MtoZ Biolabs?
- High Throughput: Capable of simultaneously identifying a large number of proteins, suitable for analyzing complex samples.
- High Sensitivity: Effectively detects low-abundance proteins, ensuring the accuracy of the analysis results.
- Reliable Results: Employs stringent quality control standards to ensure high reproducibility and comparability of the analysis results.
- Fast Delivery: Efficient analysis workflow ensures rapid delivery of experimental results and report generation.
- One-Stop Service: Provides comprehensive support throughout the entire process, from sample evaluation, data collection, to report delivery.
Applications of Large-Scale Protein Identification Service
1. Antibody Development and Engineering
Used in the antibody development process to confirm the protein composition and functionality of recombinant antibodies, ensuring product consistency.
2. Drug Target Screening
Applicable in drug development for the identification and validation of target proteins, supporting the discovery of drug targets and the study of their mechanisms of action.
3. Cell Signaling Research
Helps identify key proteins in cell signaling research and understand their role in signal transduction.
4. Biopharmaceutical Quality Assessment
Evaluates protein purity, stability, and functionality during biopharmaceutical production to ensure product quality and consistency.
Deliverables
- Comprehensive Experimental Details
- Materials, Instruments, and Methods
- Protein Identification Results Table
- Mass Spectrometry Images
- Raw Data Files
- Comprehensive Analysis Report
FAQ
Q1: What types of samples are suitable?
A1: Suitable for soluble, clarified, and high-purity protein samples, including proteins from various sources, such as recombinant proteins, fusion proteins, native proteins, antibodies, purified proteins, and peptides. Samples should avoid high salt, strong viscosity, or complex matrix interference to ensure the stability and accuracy of mass spectrometry analysis.
Q2: What is the service general workflow?
A2:
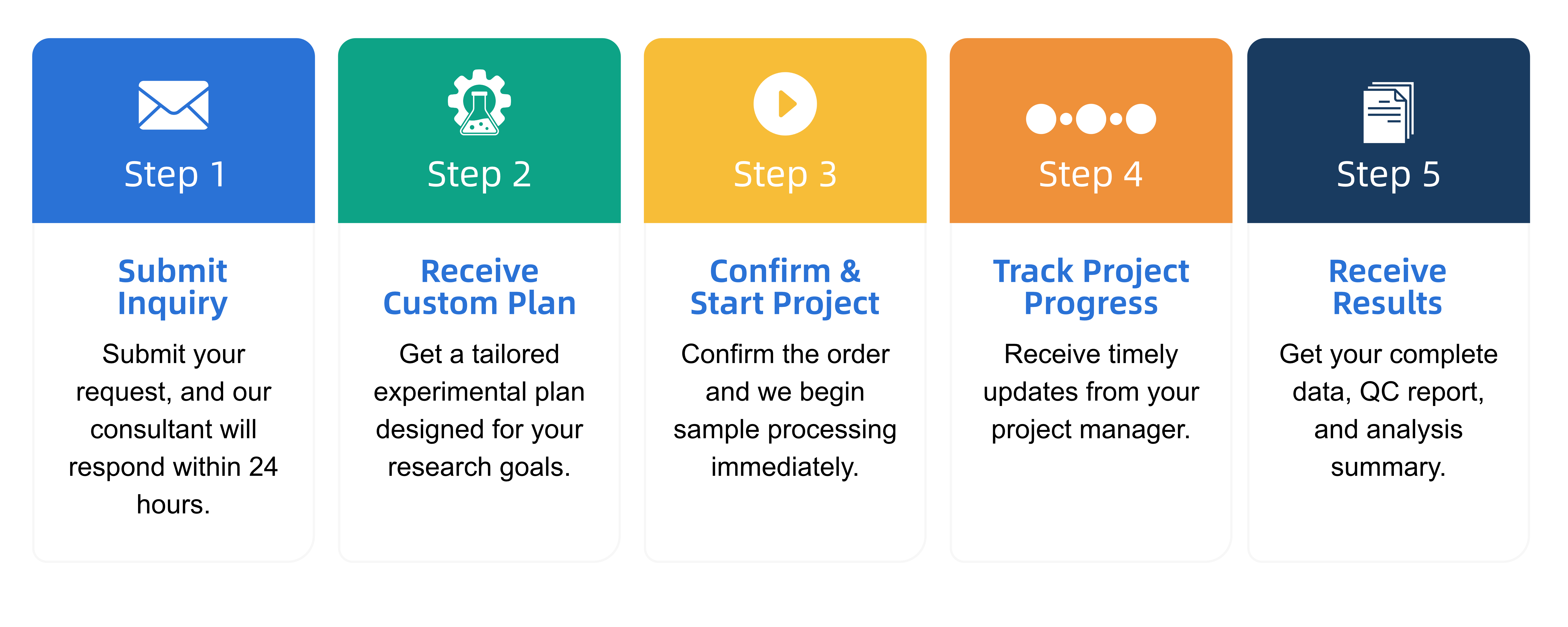
Q3: What data formats are provided?
A3: The standard data formats provided by MtoZ Biolabs include:
- Protein identification result tables (XLSX/CSV format)
- Mass spectrometry images and fragment spectra (PNG/TIFF format)
- Raw mass spectrometry data files (instrument-compatible formats)
- Comprehensive analysis reports (PDF format), including experimental methods, parameters, and identification conclusions.
If special analytical requirements exist, data formats can be customized according to project specifications.
Q4: How should I prepare my samples?
A4: The following preparation guidelines are recommended:
- Purity: Ensure the protein solution is free of precipitation and turbidity, and minimize salt, glycerol, and surfactant residues.
- Storage: Store at 4℃ for short-term and -80℃ for long-term, avoiding repeated freeze-thaw cycles.
- Transport: Use cold chain transport (ice packs/dry ice) and ensure samples are sealed to prevent leakage and avoid particle contamination.
- Additional Information: Provide sample source, concentration estimation, solvent composition, and any known modifications or expected modifications to help optimize the analysis process and result calibration.
For more information, please refer to Sample Submission Guidelines for Proteomics, Sample Submission Guidelines for Metabolomics.
Start Your Project with MtoZ Biolabs
Contact us to discuss your experimental design or request a quote. Whether you're exploring unknown sequences in proteomics or investigating key features of protein function and structure, MtoZ Biolabs can provide accurate protein identification support.
