MtoZ Biolabs provides iTRAQ-based Quantitative Proteomics Analysis Service using isobaric iTRAQ tagging combined with high-resolution LC-MS/MS to achieve precise multiplex protein quantification across multiple samples in a single experiment. This service is ideal for comparative proteomics studies such as disease versus control, multi-group designs, and time course or dose response analyses where consistent, channel-based quantitation is required.
What Is iTRAQ-Based Quantitative Proteomics?
iTRAQ-based quantitative proteomics is a multiplexed mass spectrometry strategy that labels peptides from different samples with isobaric iTRAQ tags to achieve simultaneous relative quantification. Each iTRAQ reagent contains three main components:
1. Peptide Reactive Group
- Reacts with peptide N termini and lysine side chains.
- Provides stable covalent labeling of most peptides in each sample.
2. Balance Group
- Adjusted so that the combined mass of the reporter group and balance group is the same for all tags.
- Ensures that all labeled peptides are isobaric at the precursor level and co-elute in LC-MS.
3. Reporter Ion Group
- Generates distinct reporter ions upon fragmentation during MS/MS.
- The intensity of each reporter ion reflects the relative amount of the peptide in each labeled sample.
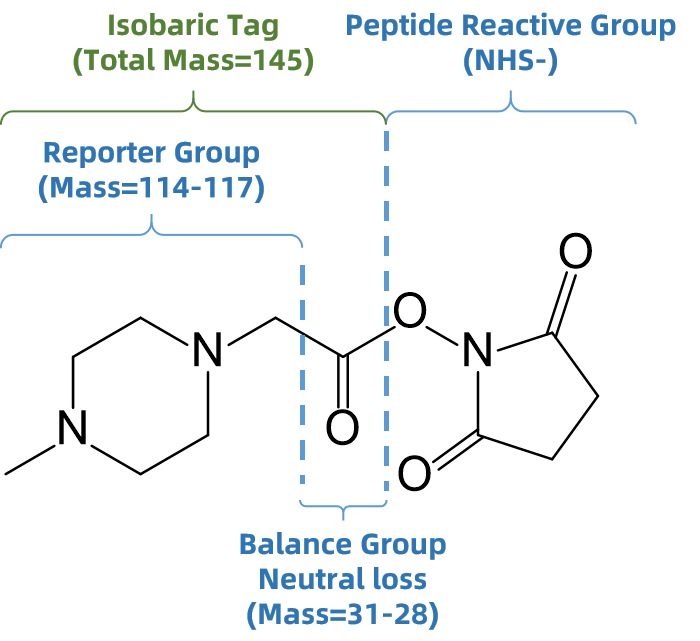
Figure 1. Structure of iTRAQ Tag
iTRAQ-Based Quantitative Proteomics Analysis Service at MtoZ Biolabs
MtoZ Biolabs offers a comprehensive iTRAQ-based Quantitative Proteomics Analysis Service built on Thermo Fisher's Q Exactive HF and Orbitrap Fusion Lumos mass spectrometry platforms coupled with Nano-LC and optimized iTRAQ workflows. You only need to define your experimental objectives and send us your samples, and we will manage the entire process, including protein extraction, digestion, iTRAQ peptide labeling, peptide separation, high-resolution LC-MS/MS acquisition, quantitative data processing, and bioinformatics interpretation tailored to your study.
In addition to iTRAQ-based quantitative proteomics, MtoZ Biolabs also provides TMT-based quantitative proteomics, SILAC-based quantitative proteomics, and label-free quantitative proteomics services to meet diverse experimental designs and analytical requirements.
Workflow of iTRAQ-Based Quantitative Proteomics Analysis Service
1. Sample Preparation and Protein Digestion
Proteins are extracted from cells, tissues, or biofluids, quantified, and enzymatically digested into peptides under standardized conditions.
2. iTRAQ Labeling and Sample Pooling
Peptides from each group are labeled with different iTRAQ reagents, checked for labeling efficiency, and then pooled for multiplexed analysis.
3. LC-MS/MS Acquisition
The combined peptide mixture is separated by Nano-LC and analyzed on high-resolution MS/MS platforms such as Q Exactive HF or Orbitrap Fusion Lumos to obtain identification and reporter ion signals.
4. Data Processing and Quantitative Analysis
Specialized software is used for peptide and protein identification, reporter ion–based ratio calculation, normalization, statistical comparison, and generation of differential protein lists with basic functional annotations.
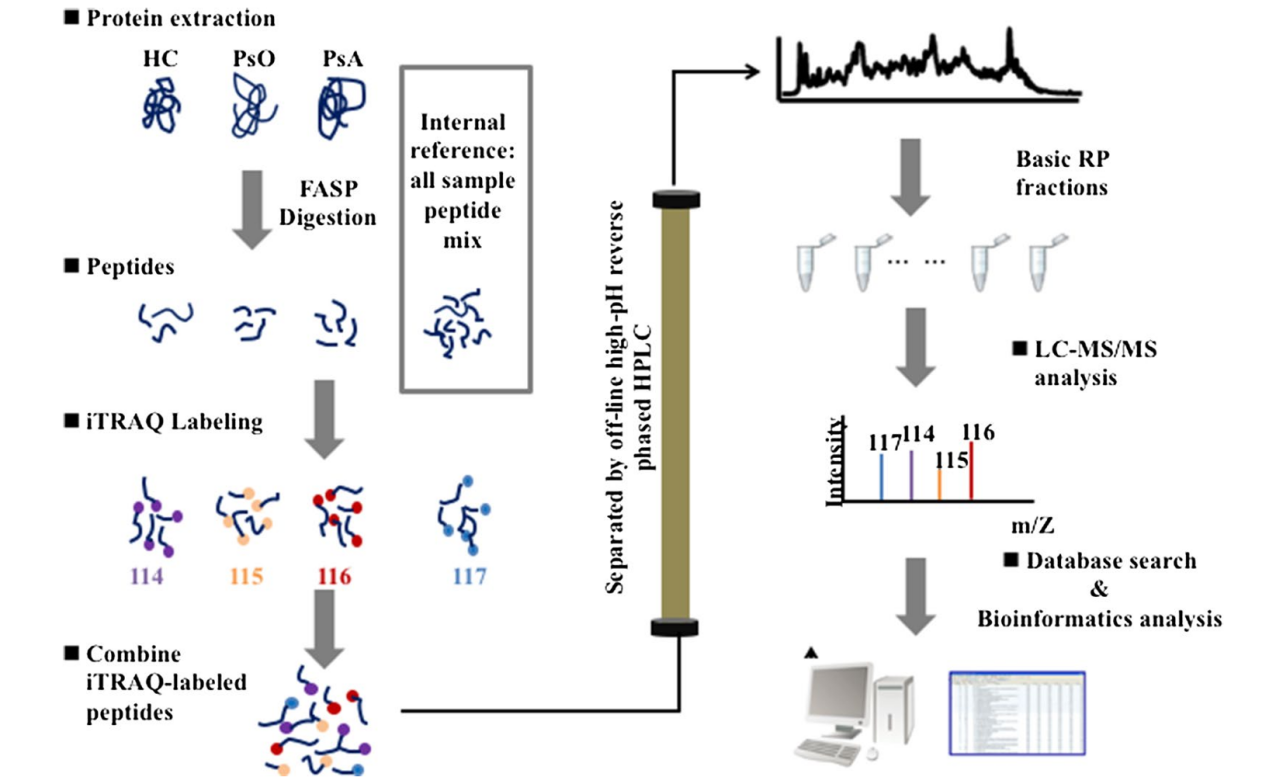
Zhu, J. et al. J Transl Med. 2021.
Figure 2. Experimental Workflow for iTRAQ Labeling Proteome Analysis
Why Choose MtoZ Biolabs
1. Stable and Reproducible Workflows
Standardized protocols for lysis, digestion, labeling, and LC-MS/MS acquisition support consistent results across projects.
2. Flexible Experimental Design
Suitable for multi-group designs, time course studies, and dose response experiments.
3. Experienced Scientific Team
Proteomics specialists provide support for study design, sample planning, and interpretation of complex quantitative data.
4. Multiplex Quantification
Analyze several samples in one run to increase throughput and reduce instrument time per sample.
5. Improved Comparability
All channels share the same chromatographic and MS conditions within a set, which reduces technical variation.
6. Deep Coverage Options
Optional fractionation and extended gradients enable detection and quantification of thousands of proteins.
Applications of iTRAQ-Based Quantitative Proteomics Analysis Service
1. Analysis of Biological Regulatory Mechanisms
Profile proteins involved in cell proliferation, differentiation, apoptosis, and other key processes to clarify their functions and regulatory pathways.
2. Signal Transduction and Pathway Studies
Monitor protein expression changes across conditions to dissect signaling cascades and cellular communication networks.
3. Biomarker Discovery and Validation
Identify and quantify candidate protein biomarkers in tissues and biofluids for early disease detection and treatment response evaluation.
4. Drug Response and Mechanism of Action
Compare proteome profiles before and after treatment to reveal drug targets, off-target effects, and pharmacodynamic responses.
5. Metabolism and Nutrition Research
Investigate proteomic changes associated with metabolic imbalance, diet interventions, or nutritional status to support mechanistic and translational studies.
FAQ
Q1: What types of samples are suitable?
We accept a wide range of protein samples suitable for iTRAQ-based quantitative proteomics, including:
-
Cell pellets from adherent or suspension cell lines
-
Primary cells from blood, bone marrow, or tissues
-
Fresh frozen tissues or biopsies
-
Biofluids such as serum, plasma, CSF, and urine
-
Preclinical animal samples for comparative or translational studies
For special sample types or low-yield materials, please contact us in advance for tailored preparation guidance.
Q2: How should I prepare my samples?
- Ensure samples are freshly prepared or snap-frozen.
- Avoid high concentrations of primary amine buffers (for example high Tris), strong detergents, and high salt that interfere with iTRAQ labeling.
- Use clean, low-binding tubes and avoid repeated freeze–thaw cycles
- Store samples at –80 °C before shipment.
- Transport samples on dry ice to maintain stability during delivery.
Detailed requirements and recommendations can be found in the Sample Submission Guidelines for Proteomics.
Q3: What is the service general workflow?
Q4: What data formats are provided?
- Raw LC-MS/MS files in instrument-specific formats
- Peptide and protein identification and iTRAQ reporter-based quantification tables in .xlsx or .csv
- Differential expression and basic functional enrichment outputs in .xlsx or .csv
- Publication-ready figures such as heatmaps, volcano plots, and clustering or PCA plots in high-resolution PNG or TIFF
- A concise PDF report summarizing workflow, QC metrics, main quantitative results, and key biological interpretations
- Additional or custom formats can be provided on request
Start Your Project with MtoZ Biolabs
Our iTRAQ-based Quantitative Proteomics Analysis Service is designed to provide more rapid, high-throughput, and cost-effective analysis, with exceptional data quality and minimal sample consumption. Free project evaluation, welcome to learn more details.
