MtoZ Biolabs provides proteomics solutions for microbial research, supporting studies on bacteria, fungi, viruses, and complex microbial communities. Our Microbial Research solutions use advanced mass spectrometry based proteomics to investigate pathogenesis, antimicrobial resistance, stress responses, and environmental impacts at the protein level.
Proteomics in Microbial Research
Microorganisms rely on tightly regulated protein networks to adapt, colonize hosts, evade immune responses, and survive in changing environments. While genetic information describes potential, microbial proteomics reveals the actual functional state of cells by measuring protein abundance, post translational modifications, secretion, and interaction networks under defined conditions.
Microbial proteomics enables researchers to:
-
Map virulence factors and host interaction proteins in pathogens
-
Characterize antimicrobial resistance mechanisms and compensatory pathways
-
Identify stress responsive proteins under heat, oxidative, osmotic, or nutrient stress
-
Track functional shifts in microbial communities under environmental or host derived pressures
By integrating high resolution LC MS/MS, quantitative proteomics, and targeted validation, MtoZ Biolabs helps you connect microbial protein changes with phenotypes such as pathogenicity, resistance, persistence, biofilm formation, and environmental functions.
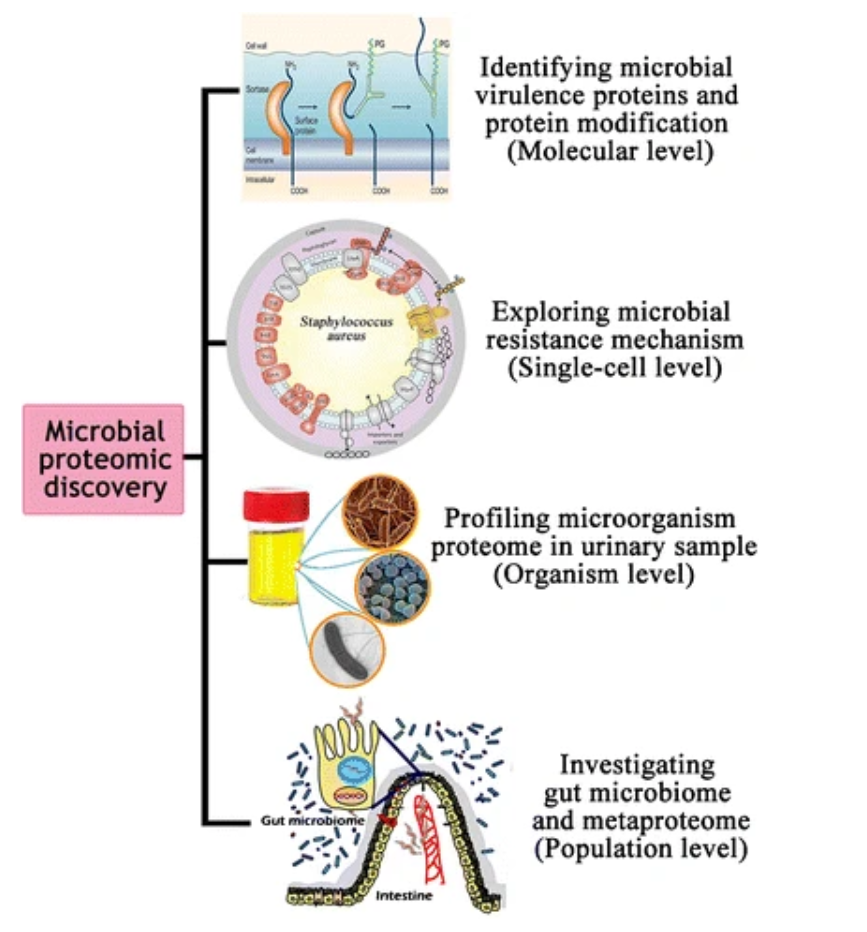
Chen, B. et al. Eur J Clin Microbiol Infect Dis. 2017.
Figure 1. Host-Microbial Pathogen Interactions from Proteomics Dissection
Service at MtoZ Biolabs
1. Pathogenesis and Virulence Mechanism Analysis
Methods
-
Global quantitative proteomics (label free, DIA, TMT based)
-
Subcellular proteomics for membrane, secretome, and surface associated proteins
-
PTM analysis for phosphorylation, acetylation, and other regulatory modifications
Service Content
-
Comparative proteome profiling of wild type vs mutant strains, or different infection stages
-
Identification of virulence associated proteins, secreted effectors, and adhesion factors
-
Mapping of altered pathways and protein networks during host interaction
Typical Applications
-
Dissecting virulence mechanisms in bacterial, fungal, or parasitic pathogens
-
Characterizing host adapted vs environmental forms of opportunistic pathogens
-
Supporting vaccine antigen selection and target prioritization
2. Antimicrobial Resistance and Drug Response Profiling
Methods
-
Quantitative proteomics for resistant vs susceptible isolates
-
Targeted proteomics (SRM PRM) for resistance markers and drug targets
-
DIA based profiling for high coverage resistance associated changes
Service Content
-
Proteome wide comparison of strains under antibiotic exposure vs controls
-
Identification of resistance related transporters, modifying enzymes, and stress proteins
-
Analysis of compensatory pathways and fitness cost adaptations
Typical Applications
-
Mechanistic studies of multidrug resistance in clinical isolates
-
Screening and verification of protein biomarkers for resistance prediction
-
Evaluating mode of action and off target effects of new antimicrobial candidates
3. Stress Response and Adaptation Protein Screening
Methods
-
Time course quantitative proteomics for stress challenge experiments
-
4D label free or DIA workflows for complex stress response profiling
-
PTM focused analysis to capture rapid regulatory events
Service Content
-
Protein expression changes under heat, cold, oxidative, osmotic, pH, or nutrient stress
-
Identification of chaperones, repair enzymes, regulators, and metabolic switches
-
Cluster analysis of co regulated proteins and stress specific signatures
Typical Applications
-
Engineering stress tolerant industrial strains for fermentation and bioprocessing
-
Understanding persistence, dormancy, and survival strategies in pathogens
-
Defining protective pathways relevant to vaccine or drug target discovery
4. Environmental Impact and Microbial Community Function
Methods
-
Metaproteomics using 4D LFQ and DIA platforms
-
Fractionation strategies for improved coverage of environmental samples
-
Functional annotation and pathway enrichment for complex communities
Service Content
-
Global functional profiling of soil, water, sludge, biofilm, and host associated microbiomes
-
Quantitative comparison of communities under different environmental conditions or treatments
-
Annotation of enzymes involved in biogeochemical cycles, degradation, and resistance
Typical Applications
-
Evaluating the effect of pollutants, antibiotics, and heavy metals on microbial functions
-
Studying biofilms in industrial, environmental, or clinical contexts
-
Supporting bioremediation projects and environmental risk assessment
Analysis Workflow
1. Sample Preparation and Collection
Microbial cells or communities are collected from sources such as water, soil, gut microbiota, or cultured strains. Depending on the matrix, samples can be:
-
Enriched and concentrated by centrifugation, membrane filtration, or ultrafiltration to isolate microbial cells
-
Processed directly from the environmental matrix when direct protein extraction is feasible
2. Community Protein Extraction
Proteins are released from microbial cells using optimized lysis buffers combined with mechanical disruption such as ultrasonication, or french pressure lysis. The extracted proteins can be used for label free or labeling based quantitative proteomics.
3. Protein and Peptide Separation
To improve identification depth and resolution, extracted proteins or their tryptic peptides are separated using:
-
One dimensional or two dimensional gel electrophoresis, with selected spots excised for in gel digestion
-
HPLC-based separation to separate complex peptide mixtures prior to MS analysis
4. Mass Spectrometry Analysis
Digested peptides are analyzed by high resolution MS and MS/MS. The instrument acquires large scale spectral data across the sample, providing the basis for comprehensive protein identification and quantification.
5. Protein Identification and Functional Interpretation
MS/MS data are processed using:
-
Database searches against microbial or metaproteomic sequence databases
-
Peptide mass fingerprinting and, when needed, de novo assisted sequencing for novel proteins
The resulting protein lists are then annotated and interpreted to reveal virulence factors, resistance mechanisms, stress response pathways, and environmental adaptation strategies in microbial systems.
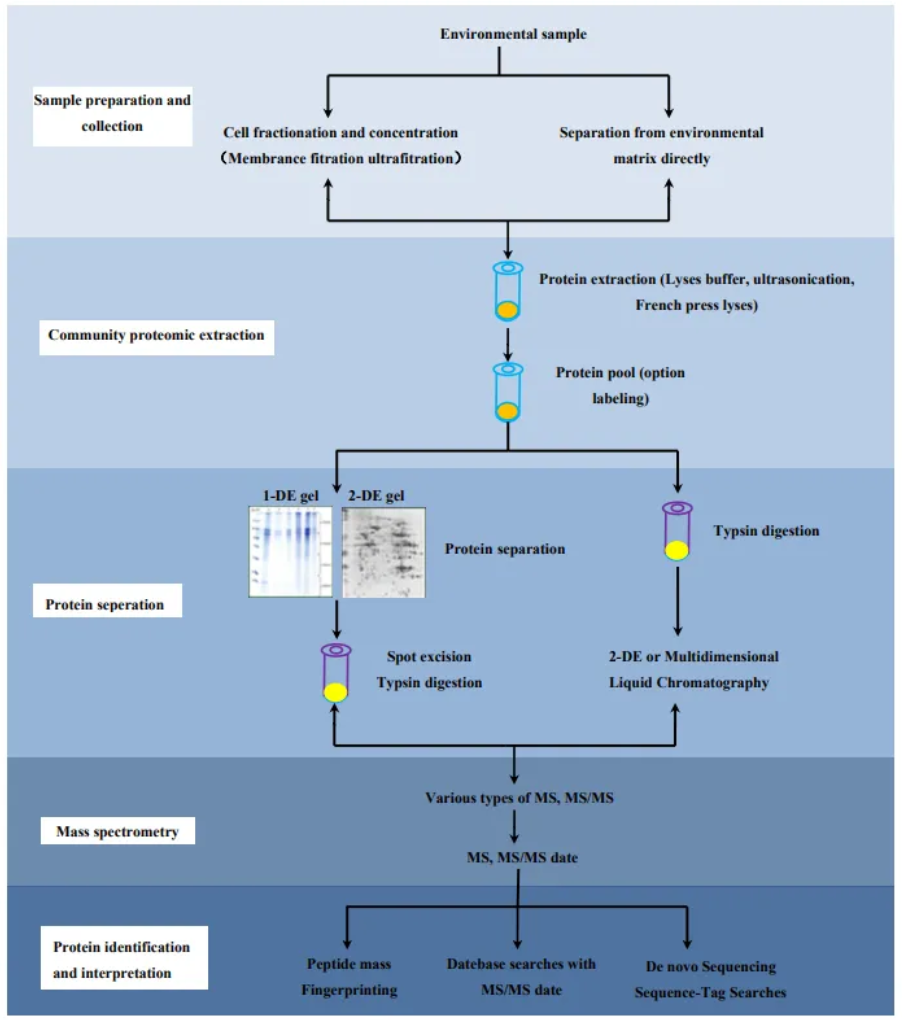
Wang, D. Z. et al. Int J Mol Sci. 2016.
Figure 2. Typical Workflow for Microorganisms Proteomics Analysis
Why Choose MtoZ Biolabs
1. Microbe Focused Expertise
Extensive experience with bacteria, fungi, archaea, and complex microbial communities, including clinical isolates, industrial strains, and environmental samples.
2. Advanced Proteomics Platforms
High resolution LC MS systems combined with DIA, 4D proteomics, and multiplexed quantitative workflows to support both discovery and targeted studies.
3. Flexible and Scalable Study Designs
Support for small exploratory projects and large cohort or time course studies with consistent pipelines and robust quality control.
4. End to End Project Support
From experimental design and sample preparation guidance to data delivery and result discussion, with a dedicated technical team for each project.
FAQ
Q1: What types of samples are suitable?
We support a wide range of microbiology-related sample types, including but not limited to:
-
Pure cultures of bacteria, fungi, yeast, and archaea
-
Mixed microbial communities (for example, soil, sediment, wastewater, marine samples)
-
Host-associated microbiota (gut, oral, skin, respiratory, plant rhizosphere)
-
Clinical and veterinary isolates or infection models
If you have other sample types, you can contact us to confirm feasibility.
Q2: How should I prepare my samples?
- Ensure samples are freshly prepared or snap-frozen.
- Avoid detergents, salts, and reducing agents that may interfere with LC-MS/MS analysis.
- Store samples at –80 °C before shipment.
- Transport samples on dry ice to maintain stability during delivery.
- provide detailed metadata (strain, growth conditions, treatments, time points)
Detailed requirements and recommendations can be found in the Sample Submission Guidelines for Proteomics.
Q3: What is the service general workflow?
Q4: What data formats are provided?
- Raw data files in instrument-specific formats (for example, .raw, .wiff, or .d)
- Processed peptide and protein identification and quantification tables in .xlsx or .csv
- Statistical and functional analysis outputs such as differential expression, pathway and GO enrichment, and taxonomic or functional annotation tables in spreadsheet formats
- Publication-ready figures, for example heatmaps, volcano plots, PCA and clustering plots, provided as high-resolution .png or .tiff
- A structured PDF report summarizing workflow, QC metrics, key results, and biological interpretation
- Additional formats or customized deliverables can be provided according to your project needs.
Start Your Project with MtoZ Biolabs
Whether you are dissecting pathogenic mechanisms, characterizing antimicrobial resistance, exploring stress responses, or profiling complex microbiomes, MtoZ Biolabs can provide a proteomics solution aligned with your project goals.
Contact us with your project objectives, and we will help you design an efficient, data-driven proteomics strategy for your microbial research.
