MtoZ Biolabs provides Global Proteomic Quantification Service to support large scale, quantitative protein profiling across diverse biological systems. Our solutions integrate multiple quantitative proteomics strategies to measure proteome wide changes in protein abundance and regulation in cells, tissues, and biofluids.
Introduction
Global proteomic quantification refers to the unbiased, large scale measurement of protein abundance and dynamic changes across conditions, time points, or treatment groups. Instead of focusing on a small predefined panel, global approaches capture thousands of proteins in a single experiment, providing a systems level view of biological states.
Proteins are the direct effectors of cellular function and key mediators of signaling, metabolism, structure, and regulation. Quantifying global protein expression and post translational modifications allows researchers to:
- Map pathways and networks that drive disease mechanisms
- Identify biomarkers associated with diagnosis, prognosis, or treatment response
- Characterize pharmacodynamic effects and off target responses
- Compare phenotypes across species, tissues, and experimental models
Global proteomic quantification is therefore central to mechanism of action studies, discovery biology, translational research, and early stage biomarker and target identification.
Global Proteomic Quantification Service at MtoZ Biolabs
1. DIA Quantitative Proteomics Analysis Service
Our DIA service provides a robust platform for protein identification and quantification, using multiple DIA strategies to achieve high sensitivity and reproducibility. With options like Direct DIA, GPF-DIA, PCT-DIA, SWATH-MS, 4D-DIA, and MSX-DIA, we offer tailored solutions for both discovery and targeted proteomics across complex biological samples.
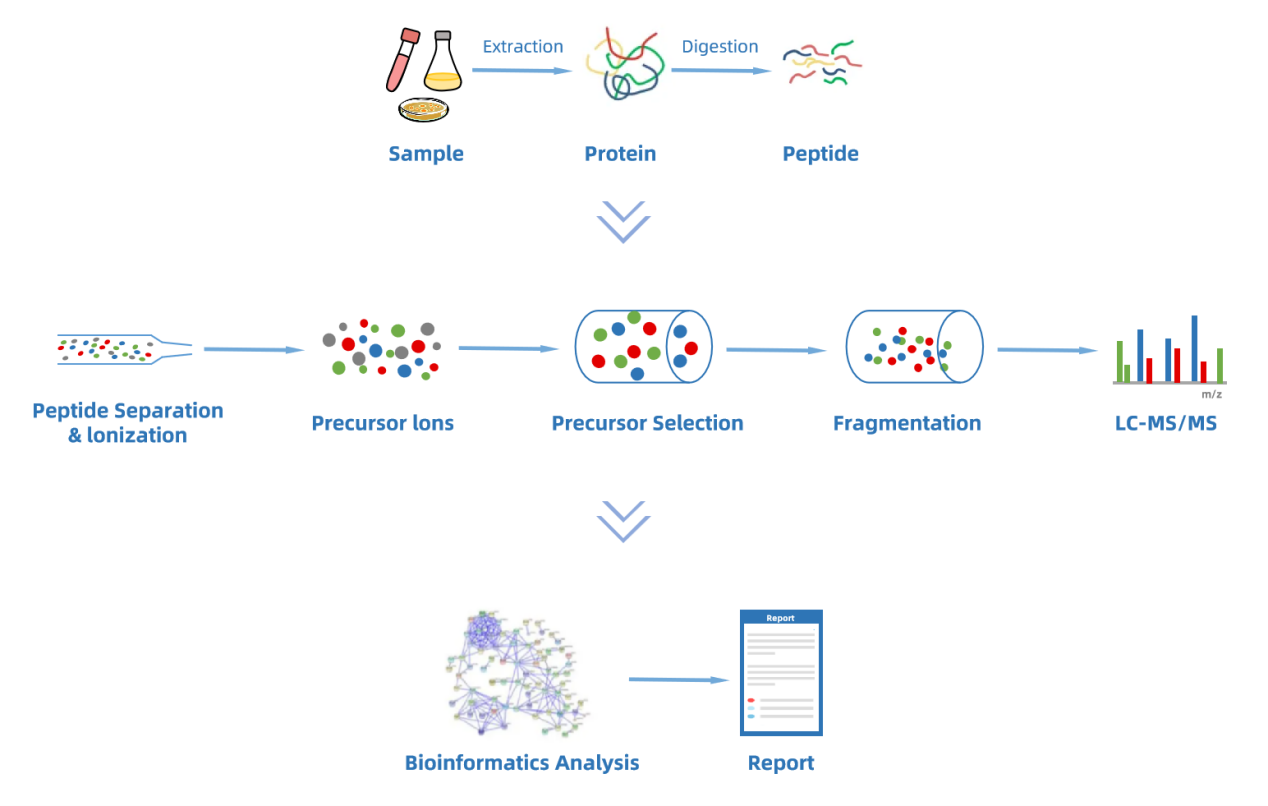
Figure 1. Workflow for DIA Quantitative Proteomics Analysis Service
2. Label-free Quantitative Proteomics Analysis Service
The label-free proteomics approach enables unbiased, direct quantification of proteins by measuring signal intensity without the need for labeling. This method is ideal for comparing protein expression across different conditions or time points in complex samples.
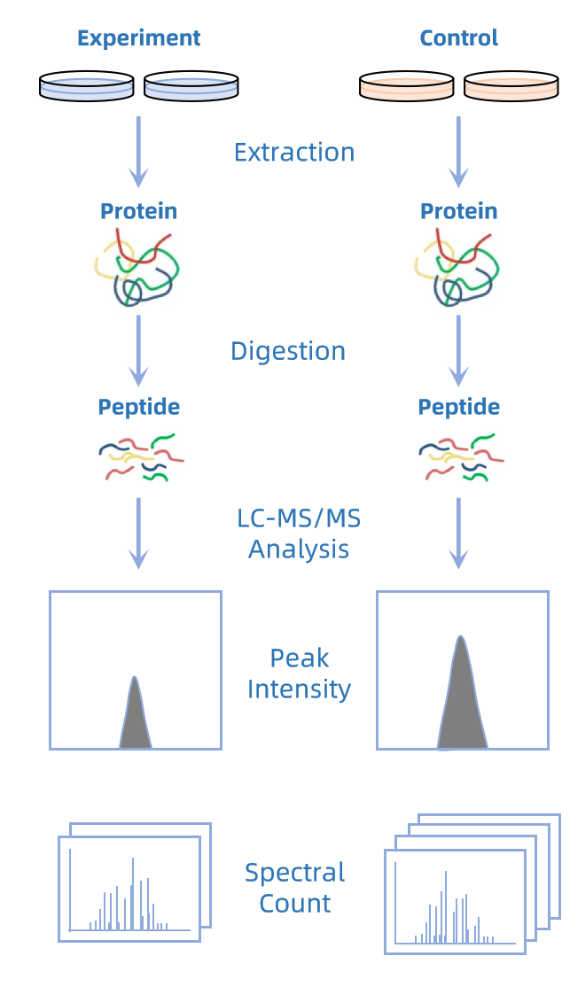
Figure 2. Principle of Label-free Quantitative Proteomics Analysis
3. 4D Label-free Quantitative Proteomics Analysis Service
Our 4D label-free service incorporates ion mobility separation, improving resolution and sensitivity, particularly for low-abundance proteins. This technology enhances the depth of analysis, making it suitable for highly complex biological samples, including clinical and preclinical applications.
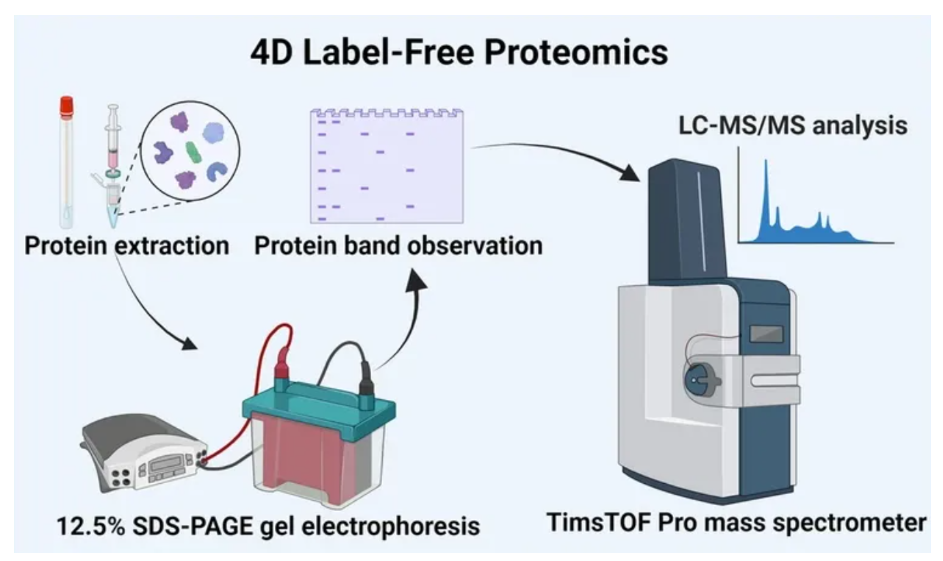
Jiang, H. et al. BMC Pregnancy and Childbirth, 2024.
Figure 3. Detection Flow of 4D Label-free Proteomics
4. TMT-based Quantitative Proteomics Analysis Service
TMT (Tandem Mass Tag) multiplexing allows the simultaneous quantification of multiple samples, improving throughput and enabling accurate comparisons. Ideal for large-scale studies, this service supports high-throughput experiments while maintaining high precision in protein quantification.
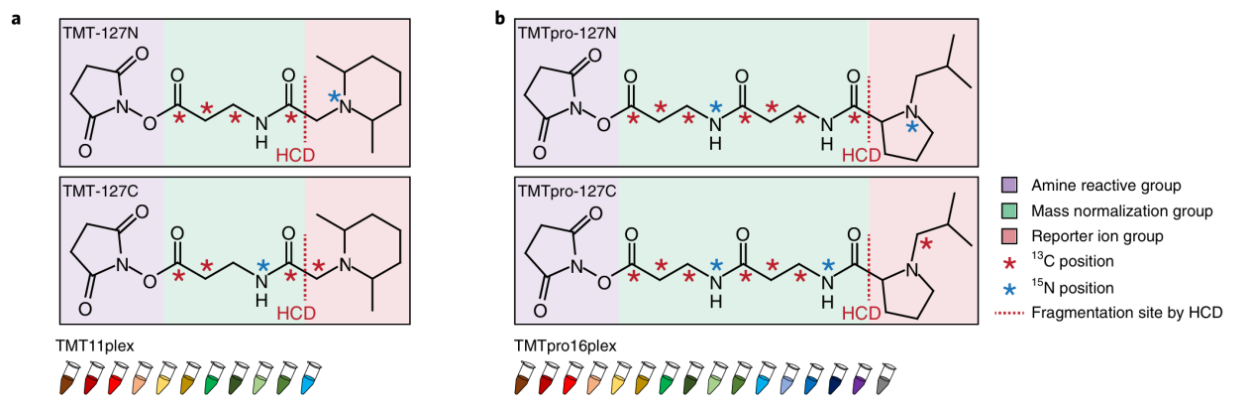
Li, J. et al. Nat Methods. 2020.
Figure 4. Chemical Structures of TMT (a) and TMTpro Reagents (b)
5. iTRAQ-based Quantitative Proteomics Analysis Service
The iTRAQ (Isobaric Tags for Relative and Absolute Quantification) technique allows the simultaneous analysis of up to 8 different samples in one experiment. This service is well-suited for comprehensive protein expression studies, providing relative quantification with high accuracy.
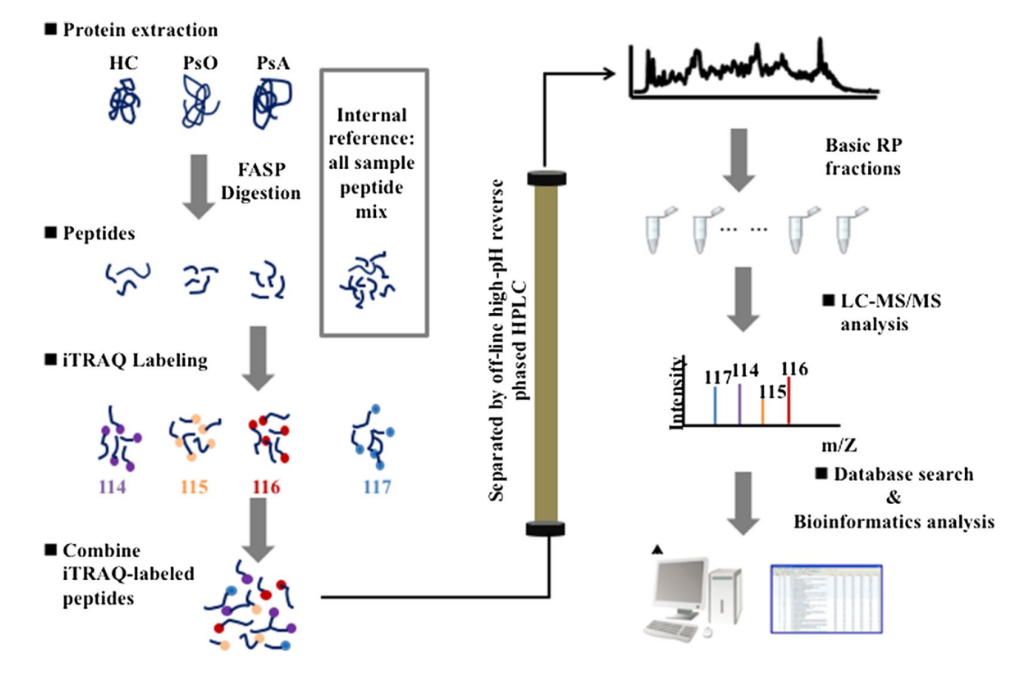
Zhu, J. et al. J Transl Med. 2021.
Figure 5. Experimental Workflow for iTRAQ Labeling Proteome Analysis
6. SILAC-based Quantitative Proteomics Analysis Service
SILAC (Stable Isotope Labeling by Amino Acids in Cell Culture) is a powerful technique for quantifying protein abundance in cultured cells. It enables precise comparison of protein expression across different experimental conditions by incorporating heavy and light isotopes into amino acids.
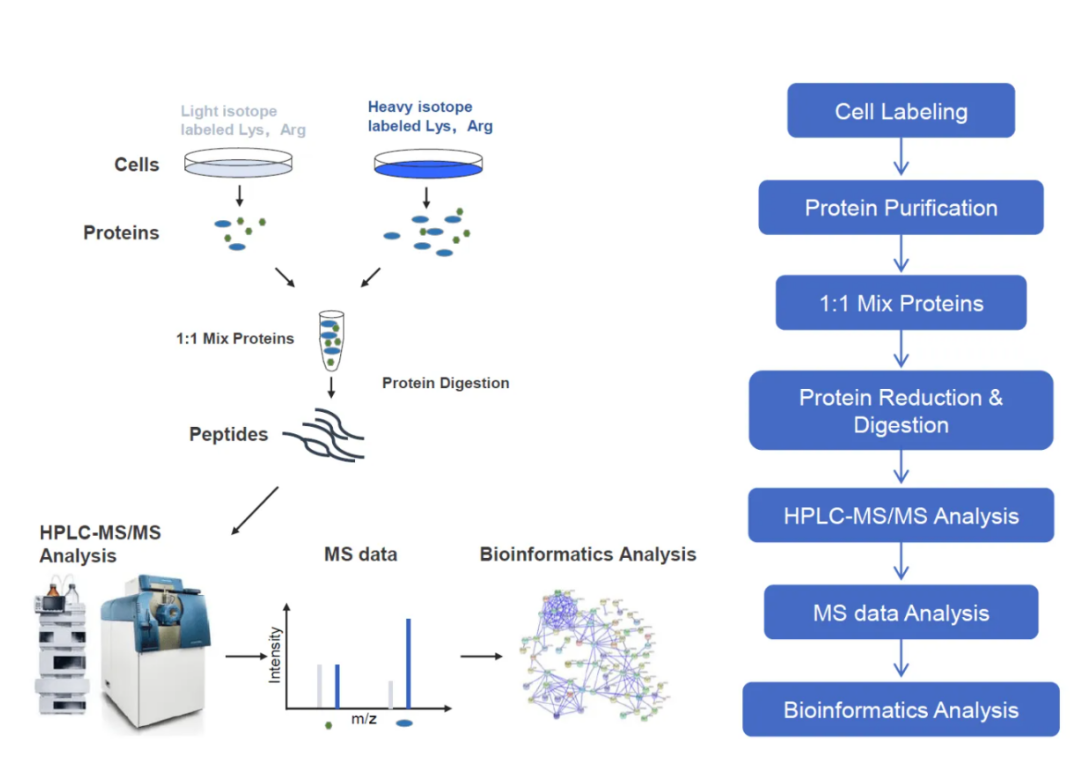
Figure 6. Workflow for SILAC-based Quantitative Proteomics Analysis Service
Why Choose MtoZ Biolabs
1. Advanced LC-MS platforms
High resolution, high sensitivity LC MS/MS systems, including ion mobility enhanced and four dimensional configurations, delivering deep proteome coverage and robust quantitation.
2. End-to-end project execution
From experimental design and sample preparation to data analysis and interpretation, we manage the complete workflow and provide clear, actionable outputs.
3. Strong quality control and reproducibility
Standardized SOPs, internal standards where appropriate, and multi level QC monitoring ensure consistent performance across runs, batches, and long term studies.
4. Flexible study scales
Support for pilot discovery projects, focused mechanistic studies, and large multi center or multi cohort projects with scalable throughput and capacity.
5. Expert scientific support
A multidisciplinary team with experience in proteomics, statistics, and biology works with you throughout the project, from planning to data review and follow up studies.
Applications of Global Proteomic Quantification Service
Global proteomic quantification at MtoZ Biolabs supports a wide range of research areas, including but not limited to:
- Disease mechanism studies: Mapping dysregulated pathways in cancer, inflammatory diseases, neurological disorders, metabolic disease, and other indications
- Biomarker discovery and prioritization: Identifying candidate markers in tissues and biofluids for early detection, prognosis, and treatment monitoring
- Drug mechanism and pharmacodynamics: Characterizing on target and off target effects, pathway modulation, and resistance mechanisms
- Toxicology and safety assessment: Profiling proteomic responses to toxicants or candidate therapeutics in relevant models
- Agricultural and plant research: Quantifying stress responses, developmental programs, and trait associated proteomes in crops and model plants
- Microbiology and host microbe interactions: Studying global proteome changes in microbes and hosts under infection, symbiosis, or environmental change
FAQ
Q1: What types of samples are suitable?
We accept a wide range of sample types for global proteomic quantification, including:
- Cell pellets
- Animal and human tissues (fresh, frozen, or FFPE)
- Plant tissues and organs
- Biofluids such as serum, plasma, CSF, urine, saliva, BALF, and others
- Microbial cultures and mixed microbial communities
- Purified proteins
If you have special or limited material, we can assess feasibility and optimize a suitable workflow.
Q2: How should I prepare my samples?
- Provide frozen cell pellets, tissue pieces, or aliquoted biofluids on dry ice or in appropriate cold-chain conditions
- Avoid repeated freeze-thaw cycles and strong detergents or chaotropes unless pre-agreed
- Clearly label all samples and provide a detailed sample list and study design
- For complex matrices or special preservatives, please contact us in advance for specific preparation guidelines
Detailed requirements and recommendations can be found in the Sample Submission Guidelines for Proteomics.
Q3: What is the service general workflow?
Q4: What data formats are provided?
- Raw LC-MS/MS data files in instrument-specific formats
- Processed identification and quantification tables for proteins and peptides in Excel or CSV formats
- Summary statistics and basic bioinformatics outputs, such as differential expression and enrichment results, in spreadsheet form
- Key visualizations, for example heatmaps, volcano plots, PCA and clustering plots, in publication-ready PNG or TIFF formats
- A structured PDF report summarizing workflow, QC metrics, main findings, and interpretation
- Additional or customized data formats can be provided according to your project requirements.
Start Your Project with MtoZ Biolabs
Whether you are exploring new mechanisms, searching for biomarkers, or quantifying treatment effects, MtoZ Biolabs Global Proteomic Quantification Service provides a flexible and comprehensive platform to support your research.
Contact us to discuss your project, and we will help you select and implement the most suitable quantitative proteomics strategy for your study.
