MtoZ Biolabs provides Single-Cell Proteomics Analysis Service to quantify proteins at true single-cell resolution, helping you dissect cellular heterogeneity in cancer, immunology, neuroscience, and developmental biology. Using ultra-sensitive LC-MS based workflows optimized for low-input samples, we deliver robust protein expression profiles and functional insights from rare or complex cell populations.
What Is Single-Cell Proteomics?
Single-cell proteomics focuses on measuring the protein composition and abundance of individual cells rather than bulk populations. While single-cell genomics and transcriptomics reveal genetic variation and RNA expression, proteins are the direct executors of cellular function, integrating transcriptional programs, post-translational modifications, and environmental signals. Single-cell proteomics therefore offers a more proximal readout of phenotype and functional state.
In many biological and disease contexts, bulk measurements mask critical subpopulations. Tumor samples contain mixtures of malignant cells, stromal cells, and infiltrating immune cells. Immune responses involve transient activation states that can only be captured at the individual cell level. Single-cell proteomics enables you to identify rare cell types, resolve functional subclusters, and understand how protein networks vary from cell to cell within the same tissue or condition.
Single-Cell Proteomics Analysis Service at MtoZ Biolabs
MtoZ Biolabs has built a dedicated single cell proteomics platform that integrates ultra sensitive sample preparation with high-resolution Orbitrap based mass spectrometry and ion mobility enhancement such as FAIMS to improve signal quality and detection of low abundance proteins. Single cells obtained by FACS, microfluidics, or pre isolated suspensions are gently lysed, digested, and prepared in miniaturized formats to minimize loss. We support TMT based multiplexing to analyze many single cells in parallel, increasing throughput while maintaining quantitative consistency across conditions.
Workflow of Single-Cell Proteomics Analysis Service
1. Sample Collection and Single-Cell Isolation
Single-cell suspensions are prepared from cultured cells, tissues, or body fluids, and individual cells are isolated using FACS, microfluidic platforms, or microdissectio.
2. Cell Lysis and Protein Digestion
Isolated single cells or low-input samples are lysed in low-volume, MS-compatible buffers, followed by reduction, alkylation, and enzymatic digestion (typically trypsin) to generate peptides.
3. Protein Identification and Quantification
Peptides are separated by nano-LC and analyzed on high-sensitivity LC-MS/MS platforms using single-cell optimized acquisition strategies. Dedicated software is then used to identify peptides and infer proteins, build quantitative matrices, and control FDR across cells and conditions.
4. Bioinformatics and Biological Interpretation
Processed single-cell protein abundance data are normalized and used for clustering, dimensionality reduction, differential abundance analysis, and pathway or network enrichment, providing clear, biologically meaningful insights that align with your study goals.
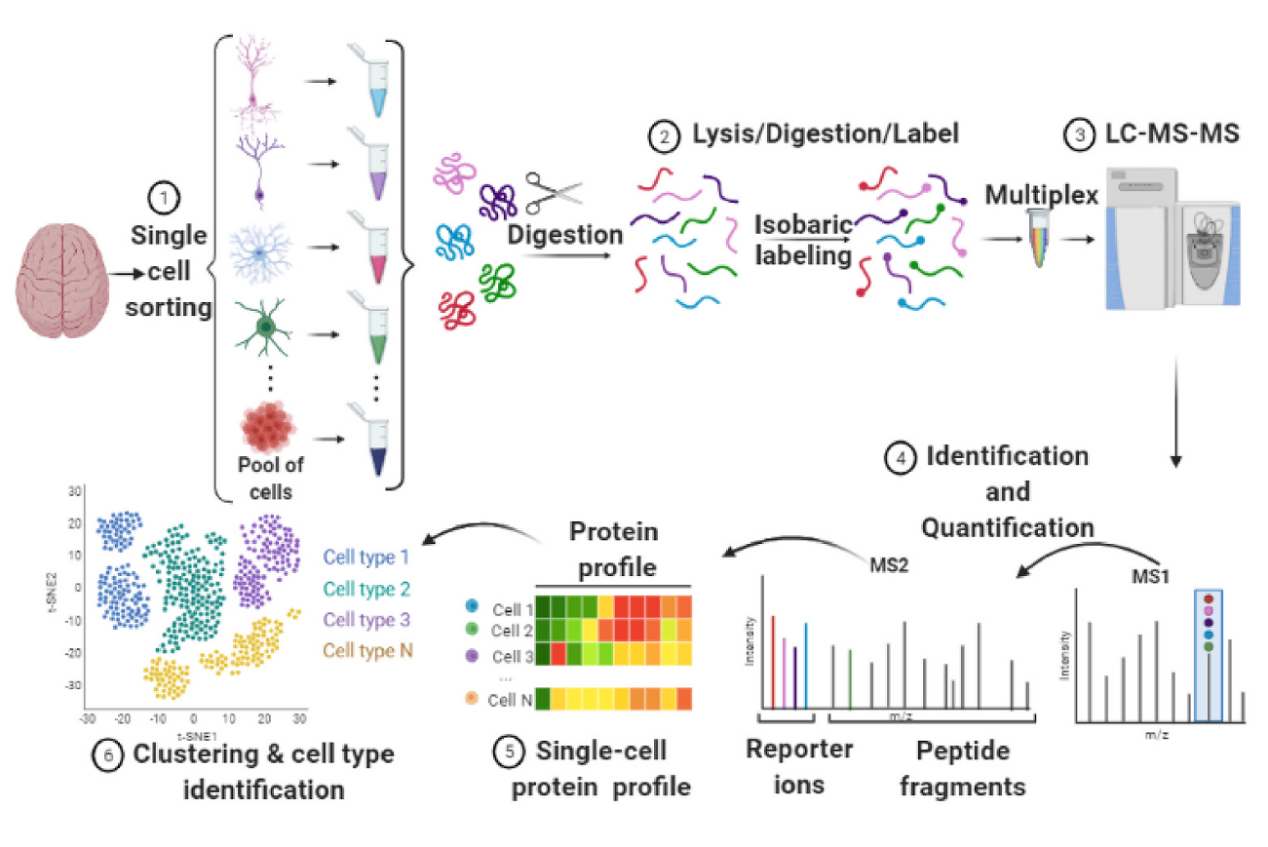
Goto-Silva, L. et al. Biochim Biophys Acta Proteins Proteom. 2021.
Figure 1. Pipeline for Single-Cell Proteomics of the Brain
Why Choose MtoZ Biolabs
1. Professional Multidisciplinary Team
Experienced proteomics scientists and bioinformaticians who understand single-cell experimental design, data challenges, and biological interpretation.
2. Customized Project Design
Flexible workflows tailored to your sample types, cell numbers, and research questions, from pilot feasibility studies to large-scale projects.
3. High Efficiency and High Data Quality
Standardized SOPs, strict QC, and optimized acquisition methods that deliver reproducible, publication-ready datasets with minimal missing values.
4. Excellent Customer Support
One-on-one scientific communication throughout the project, from study planning to result review, with clear reports and responsive technical support.
Applications of Single-Cell Proteomics Analysis Service
1. Cancer Research and Tumor Heterogeneity
Dissect subclonal diversity, drug resistance mechanisms, and tumor microenvironment at single-cell resolution.
2. Immunology and Inflammation
Profile activation states of T cells, B cells, NK cells, and myeloid cells to study immune responses, tolerance, and immunotherapy effects.
3. Neuroscience and Neurodegeneration
Characterize neurons and glial cell subtypes, synaptic protein changes, and early proteomic alterations in brain disorders.
4. Stem Cell and Developmental Biology
Track protein level changes during differentiation, lineage commitment, and reprogramming in stem and progenitor cells.
5. Precision and Translational Medicine
Analyze patient derived cells to identify clinically relevant cell states, stratify patients, and support biomarker and target discovery.
6. Host-Pathogen and Infection Studies
Investigate cell type specific host responses to viral, bacterial, or parasitic infection at the protein network level.
FAQ
Q1: What types of samples are suitable?
We support a variety of sample types, including
-
Cultured cell
-
Primary tissues
-
PBMCs or blood cells
-
Plant Cells / protoplasts
-
Enriched immune cell subsets or rare cell populations
If you have special sample types or low-yield materials, we can assess feasibility based on your project goals.
Q2: How should I prepare my samples?
- Ensure samples are freshly prepared and snap-frozen.
- Avoid harsh fixatives and high concentrations of detergents that interfere with MS.
- Store samples at –80 °C before shipment and avoid repeated freeze-thaw cycles.
- Transport samples on dry ice to maintain stability during delivery.
For more information, please refer to Sample Submission Guidelines for Proteomics.
Q3: What is the service general workflow?
Q4: What data formats are provided?
- Raw data files in instrument-specific formats (for example .raw or .d)
- Peptide and protein identification and quantitative matrices in .xlsx or .csv
- Statistical and bioinformatics result tables, including clustering and differential expression, in spreadsheet formats
- Key visualizations such as heatmaps, PCA or UMAP plots, and cluster maps in high-resolution .png or .tiff
- A concise PDF report summarizing workflow, QC metrics, main results, and biological interpretation
- Additional formats or customized deliverables can be provided upon request.
Start Your Project with MtoZ Biolabs
If you are ready to move beyond bulk measurements and explore protein level heterogeneity at the single-cell scale, MtoZ Biolabs is ready to support you.
Contact MtoZ Biolabs to discuss your project objectives, sample options, and timelines. We will help you define a single-cell proteomics strategy that fits your scientific goals and delivers data you can trust.
