MtoZ Biolabs provides a 4D-LFQ Metaproteomics Service for high-resolution, label-free quantitative analysis of complex microbial communities from environmental and host-associated samples. By integrating robust 4D LC-MS workflows with metaproteomics-optimized bioinformatics pipelines, we enable reliable protein quantification and confident functional interpretation from highly complex microbiome samples. This service supports the transition from mixed community proteomes to functionally resolved protein profiles, taxonomy-linked protein assignments, and pathway-level insights, facilitating investigations into microbial functional organization, community-level regulation, and microbiome-associated phenotypes in diverse biological systems.
What Is 4D-LFQ Metaproteomics?
4D-LFQ metaproteomics is a mass spectrometry based approach that profiles and quantifies proteins from entire microbial communities and their host environment in a single experiment. Instead of focusing on isolated strains, it characterizes the collective proteome of bacteria, archaea, fungi, viruses, and host cells present in complex samples such as stool, soil, wastewater, biofilms, or mucosal tissues. By measuring the proteins that are actually present and active, 4D-LFQ metaproteomics provides a direct functional view of community metabolism, signaling, and interaction with the host or environment.
The "4D" component refers to the four analytical dimensions captured by modern high resolution LC-MS systems: retention time, mass to charge ratio, signal intensity, and an additional ion mobility or collision cross section dimension. Ion mobility separates peptides in the gas phase according to their size and shape, improving the resolution of coeluting species and reducing signal interference in highly complex mixtures.
LFQ, or label free quantification, uses the intensity or integrated peak area of peptide signals across runs to compare protein abundance between conditions without the need for isotopic or isobaric labels. When combined with 4D separation, LFQ enables deep and reproducible quantification of metaproteomes, including low abundance enzymes, transporters, and regulatory proteins that are critical for understanding microbial activity, community structure, and functional shifts under different biological or environmental conditions.
4D-LFQ Metaproteomics Service at MtoZ Biolabs
MtoZ Biolabs offers an end-to-end 4D-LFQ Metaproteomics Service that connects advanced 4D LC-MS analysis with microbiome optimized workflows and bioinformatics pipelines. The service is designed to support discovery driven research, hypothesis testing, and translational projects in microbiome and host microbiome biology.
You only need to define your project goals and send us your samples. MtoZ Biolabs manages protein extraction from complex matrices, digestion, LC-MS analysis, label free quantification, and metaproteomics bioinformatics, delivering a complete dataset ready for biological interpretation.
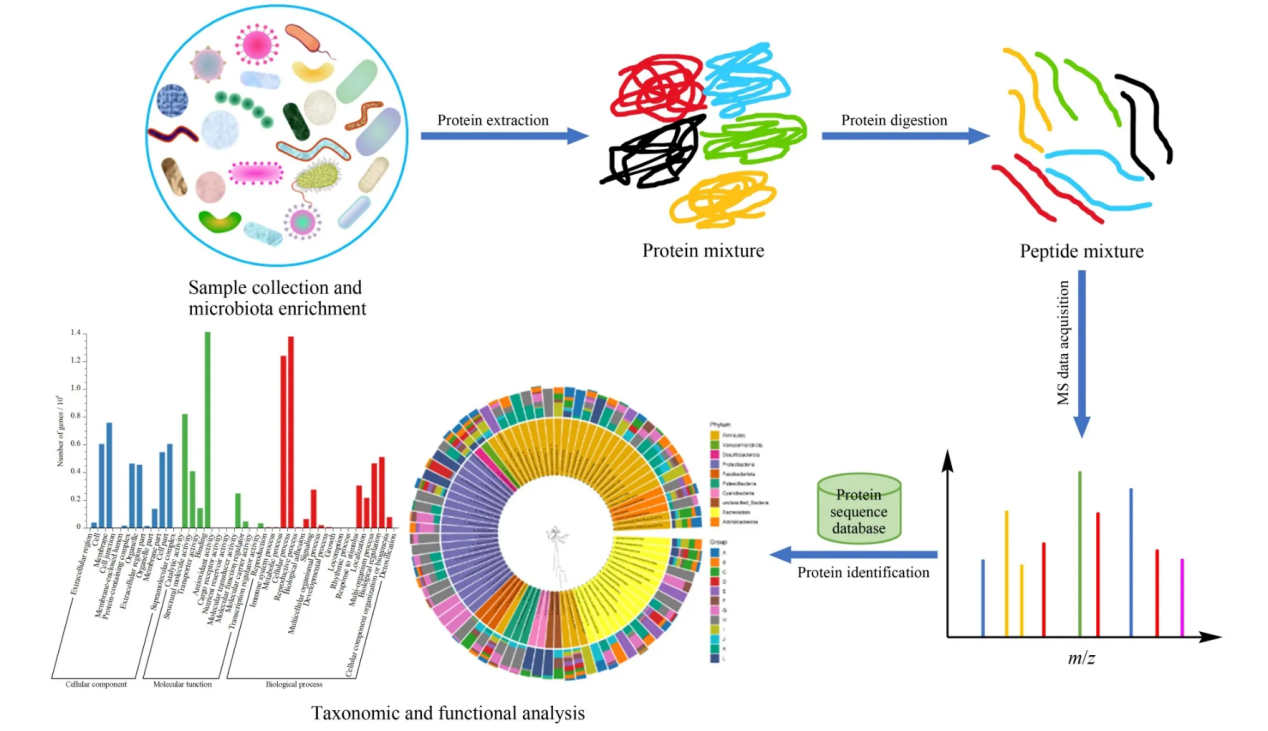
Wu, E. H. et al. Se Pu. 2024.
Figure 1. Standard Flow Chart of Metaproteomic Detection
Why Choose MtoZ Biolabs
- Advanced 4D-LFQ Metaproteomics workflows for higher sensitivity, deeper coverage, and improved separation in highly complex community samples.
- Metaproteomics optimized sample prep, acquisition, and analysis pipelines designed specifically for mixed microbial and host proteomes.
- Flexible, customizable projects with tailored databases, analysis depth, and reporting aligned with your biological questions.
- Comprehensive bioinformatics, including taxonomic and functional annotation, pathway analysis, and clear result summaries.
- Dedicated proteomics experts providing study design support, troubleshooting, and responsive communication throughout the project.
Applications of 4D-LFQ Metaproteomics Service
4D-LFQ Metaproteomics Service at MtoZ Biolabs supports multiple research areas:
1. Gut Microbiome and Human Health
-
Functional profiling of intestinal microbiota in metabolic, inflammatory, and neurological conditions
-
Evaluation of diet, probiotics, prebiotics, and therapeutics on microbiome activity
2. Environmental and Soil Microbiology
-
Analysis of soil and sediment communities involved in carbon, nitrogen, and sulfur cycling
-
Assessment of pollutant degradation, bioremediation, and ecosystem responses to environmental change
3. Industrial and Bioprocess Microbiology
-
Functional monitoring of microbial consortia in bioreactors, fermentation processes, and wastewater treatment
-
Process optimization by tracking key enzymes and metabolic pathways
4. Host Associated Microbiomes
-
Metaproteomics of skin, oral, respiratory, or plant associated microbiomes
-
Understanding host microbiome interactions and immune modulation
5. Infectious Disease Research
-
Investigation of pathogen–commensal–host protein networks in infection models
-
Detection of virulence factors, resistance proteins, and immune related signatures
FAQ
Q1: What types of samples are suitable?
We accept a wide range of microbiome and host associated samples, including but not limited to:
-
Feces and intestinal contents
-
Oral, nasal, skin, or mucosal swabs
-
Soil, sediment, sludge, and wastewater
-
Biofilms and fermentation broth
-
Environmental filters and pellets
-
Host tissues or tissue homogenates containing microbial communities
If you have other sample types, you can contact us to confirm compatibility.
Q2: How should I prepare my samples?
To ensure high quality 4D-LFQ metaproteomics data, we recommend:
- Collect samples using clean, low protein binding tubes and tools
- Snap freeze samples in liquid nitrogen or on dry ice as soon as possible
- Avoid strong detergents, high salt, chelators, and organic solvents if you perform any pre processing
- Do not add preservatives that may interfere with protein extraction or LC-MS analysis
- Store samples at -80 °C until shipment
- Ship samples on dry ice with clear labels and a sample information sheet
Detailed requirements and recommendations can be found in the Sample Submission Guidelines for Proteomics.
Q3: What is the service general workflow?
Q4: What data formats are provided?
- Raw data files in instrument specific formats
- Processed peptide and protein abundance tables in Excel or CSV
- Taxonomic and functional annotation results, including pathway and enrichment outputs, in spreadsheet formats
- Key visualizations such as heatmaps, volcano plots, clustering and PCA in high resolution PNG or TIFF
- A structured PDF report summarizing workflow, QC metrics, main findings, and biological interpretation
- Additional file formats or custom reports can be provided on request.
Start Your Project with MtoZ Biolabs
Contact us to discuss your experimental design or request a quote. Our technical specialists are available to provide a free business assessment.
