MtoZ Biolabs provides Plant Targeted Proteomics Service based on targeted mass spectrometry strategies, including Multiple Reaction Monitoring (MRM), Selected Reaction Monitoring (SRM), and Parallel Reaction Monitoring (PRM). This service is designed for accurate and reproducible quantification of predefined plant proteins across diverse biological samples and experimental conditions.
Unlike discovery-based proteomics, targeted proteomics selectively monitors predefined peptides and ion signals during mass spectrometry acquisition. This targeted data collection strategy significantly improves analytical sensitivity, quantitative precision, and reproducibility, making it particularly suitable for hypothesis-driven studies, pathway-focused research, and large sample cohorts in plant biology.
Technical Principles
Plant targeted proteomics is based on predefined peptide targets and selective ion monitoring during mass spectrometry acquisition. By focusing on specific peptides derived from proteins of interest, this approach enables accurate, sensitive, and reproducible quantification in complex plant matrices.
MRM and SRM are typically performed on triple quadrupole mass spectrometers, where both precursor ions and their corresponding fragment ions are selectively monitored using predefined ion transitions. This highly targeted data acquisition strategy provides excellent quantitative precision, wide dynamic range, and strong reproducibility, making it well-suited for large sample cohorts and low-abundance protein measurement.
PRM applies targeted precursor ion selection combined with high-resolution fragment ion detection. Instead of monitoring individual transitions, PRM records the full set of fragment ions generated from a selected precursor with high mass accuracy. This enhances selectivity and reduces interference, which is particularly advantageous for complex plant samples containing abundant background components.
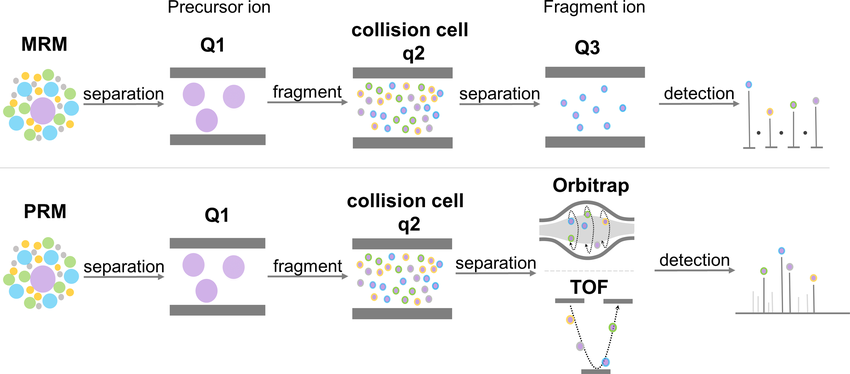
Yin, H. et al. Mass Spectrometry Reviews. 2022.
Figure 1. Schematic Diagram of the MRM and PRM
Together, MRM/SRM and PRM enable robust, antibody-independent targeted proteomics analysis, supporting reliable quantification of predefined plant proteins across different tissues, treatments, and developmental stages.
Plant Targeted Proteomics Service at MtoZ Biolabs
MtoZ Biolabs offers Plant Targeted Proteomics Service using established targeted mass spectrometry platforms and validated analytical strategies.
Our service capabilities include:
- MRM/SRM-based targeted protein quantification
- PRM-based high-resolution targeted proteomics
- Relative or absolute quantification using internal or isotope-labeled standards
- Targeted analysis of defined protein panels involved in metabolic or signaling pathways
Each assay is designed with careful peptide selection and method optimization to ensure stable performance and reliable quantitative output.
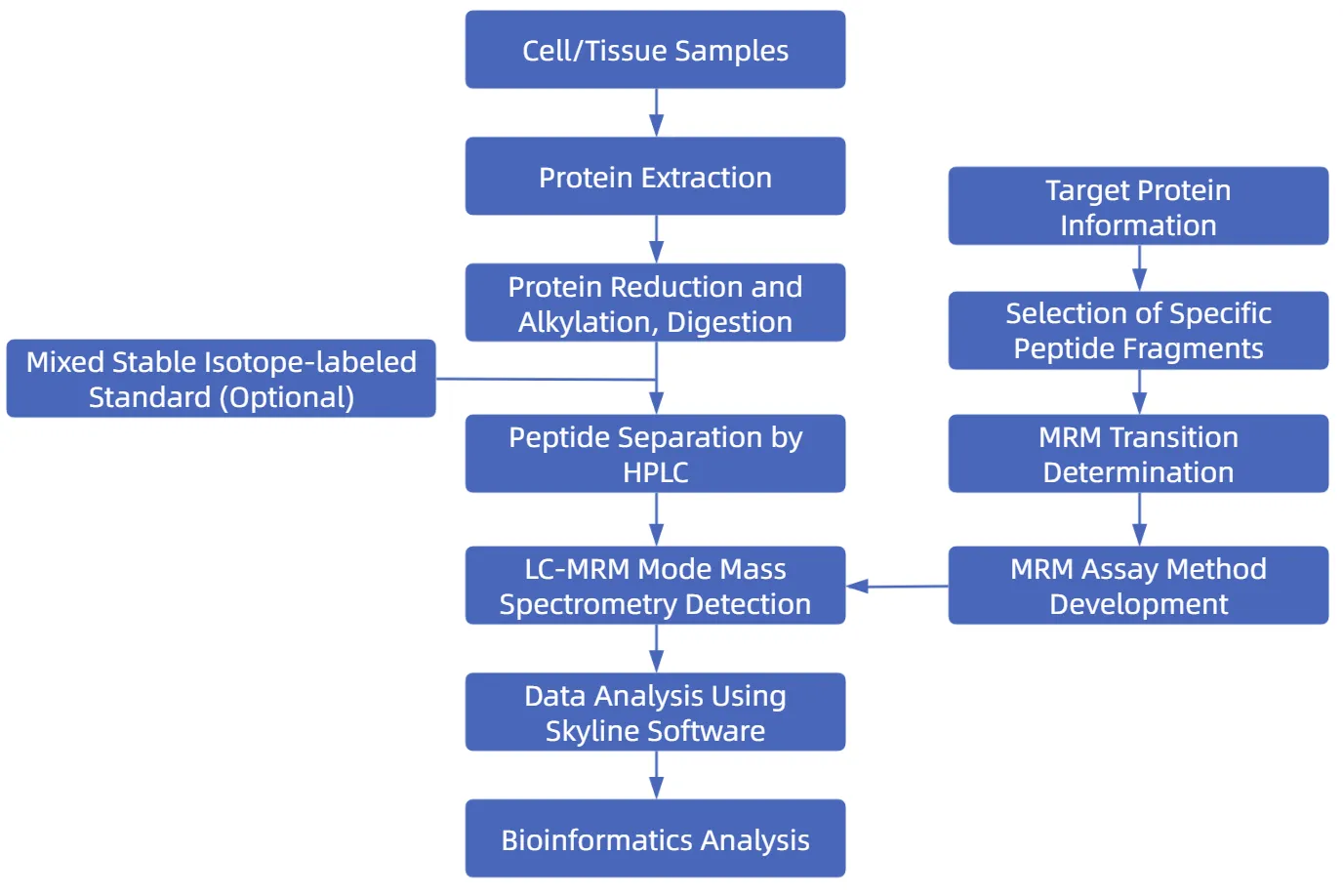
Figure 2. Workflow of MRM Proteomics
Why Choose MtoZ Biolabs?
1. Advanced Analysis Platform
Established MRM, SRM, and PRM platforms ensure accurate, sensitive, and reproducible protein quantification.
2. Plant-Oriented Analytical Design
Workflows are optimized to address the biochemical complexity and matrix effects inherent to plant samples.
3. High Sensitivity and Quantitative Precision
Targeted ion monitoring enables reliable detection of low-abundance proteins in complex plant proteomes.
4. Antibody-Independent Detection
MRM and PRM approaches eliminate reliance on antibodies, expanding applicability and avoiding antibody specificity limitations.
5. Customized Experimental Strategy
Flexible assay design and data analysis are tailored to specific research objectives and biological questions.
6. One-Time-Charge
Our pricing is transparent, no hidden fees or additional costs.
Start Your Project with MtoZ Biolabs
MtoZ Biolabs delivers Plant Targeted Proteomics Service using MRM, SRM, and PRM strategies to support precise and reproducible protein quantification in hypothesis-driven plant research. Contact us to discuss your study design or request a customized targeted proteomics solution.
FAQ
Q1: What types of samples are suitable?
This service supports a wide range of plant samples, including leaves, roots, stems, seedlings, flowers, and seeds, as well as plant cell cultures, suspension cells, and subcellular fractions. Purified plant proteins and enriched protein extracts are also compatible. There is no restriction on plant species.
Q2: How should I prepare my samples?
Samples should be prepared under conditions that preserve protein integrity and ensure consistency across experimental groups. Avoid repeated freeze-thaw cycles and excessive handling. Samples should be kept cold during processing and shipped on dry ice when applicable.
For more information, please refer to Sample Submission Guidelines for Proteomics. If you are unsure about sample preparation, our team is available for consultation and can guide you through the process to ensure optimal results.
Q3: What is the service general workflow?
Q4: What data formats are provided?
Deliverables include raw LC-MS/MS data files, quantitative result tables in Excel or CSV format, and a PDF report summarizing analytical methods, quality control, and quantitative findings. Figures and chromatograms are provided in PNG or TIFF format. Customized data outputs can be provided upon request to meet specific research or publication requirements.
Related Services
